Pymolshortcuts
The repository pymolschortucts contains the a collection of shortcuts that are loaded on startup of PyMOL. These shortcuts enable websearches from within PyMOL as well as many other convienent functions that make work in PyMOL more productive..
Install / Use
/learn @MooersLab/PymolshortcutsREADME
pymolshortcuts
This is the repository for pymolschortucts.py. This file contains 215 functions in 25 categories mapped to short names that work like aliases (the table below is incomplete). These shortcuts include many functions that make working in PyMOL more productive and fun! Some shortcuts save the many hours of work required to assemble a new script file while other shortcuts save only a few minutes but lower motivational barriers. You do not need to understand Python to be able to run the script.
A paper about this project was published in Protien Science in January 2020. Access to the PDF is open; you do not need a subscription to Protein Science to download it.
To cite the software, use the following citation if you use BibTeX.
@Article{Mooers2020ShortcutsForFasterImageCreationInPyMOL,
author = {Mooers, Blaine HM},
title = {Shortcuts for faster image creation in PyMOL},
number = {1},
pages = {268--276},
volume = {29},
file = {:Mooers2020ShortcutsForFasterImageCreationInPyMOL.pdf:PDF},
url = {https://onlinelibrary.wiley.com/doi/pdfdirect/10.1002/pro.3781},
journal = {Protein Science},
publisher = {Wiley Online Library},
doi = {10.1002/pro.3781},
year = {2020},
}
The Google Scholar generated ENDNOTE file (scholar. new) is above with the other files.
Quick overview
To get a quick overview of this project, scroll down through this README.md file and watch the animations. To learn more, see the PDFs in the slideshow folder:
- mooers23jul2019ACAPyMOLshortcuts.pdf (20-minute talk at the American Crystallographic Association annual meeting in Covington, KY)
- mooers18oct2019RNAcornbeltPyMOLShortcuts4RNA.pdf (9-minute talk at the Cornbelt RNA meeting at the U of Missouri in Columbia, Missouri)
- mooers26Oct2019SWTCCPyMOLshortcuts.pdf (20-minute talk, Southwest Theoretical and Computational Chemistry (SWTCC) annual meeting at the U of Oklahoma in Norman, OK)
The animation below demonstrates the photorealistic effect from the ambient occlusion shortcut applied to a protein by entering two letters, AO. This shortcut executes 16 lines of code with the entry of two letters!
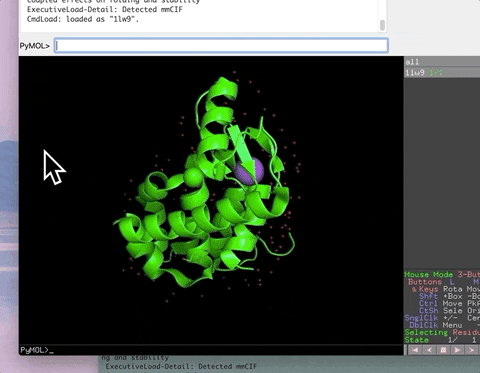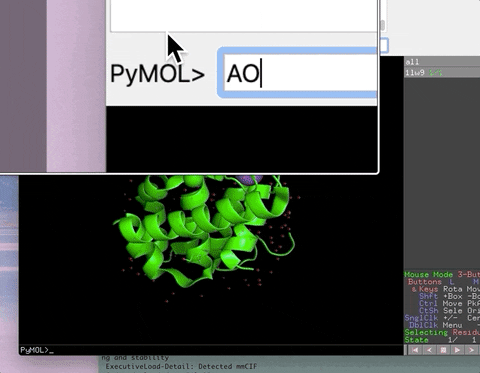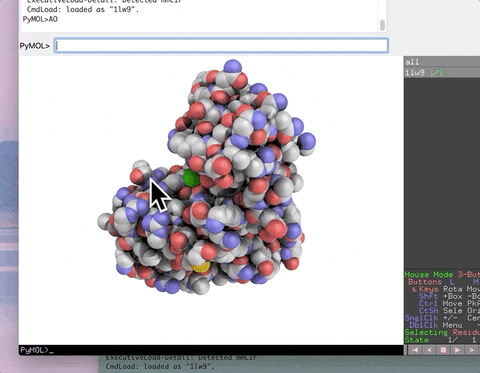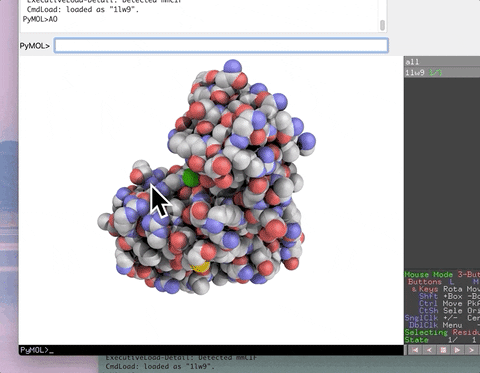
The animation below demonstrates the submission of two shortcuts, sc311 and sc113, separated by a semicolon on the command line. They generate symmetry mates that fill five unit cells. The unit cells form an L shape. Each symmetry mate appears as a separate object in the menu in the panel to the right of the vv. These can be removed by entering the shortcut rmsc.
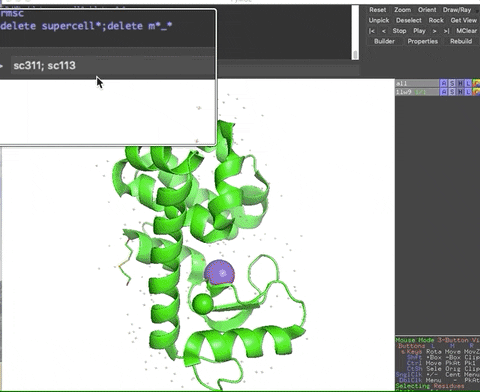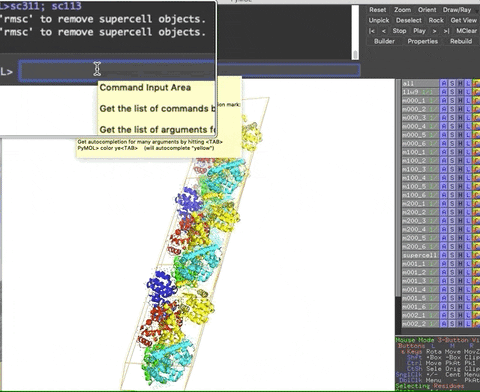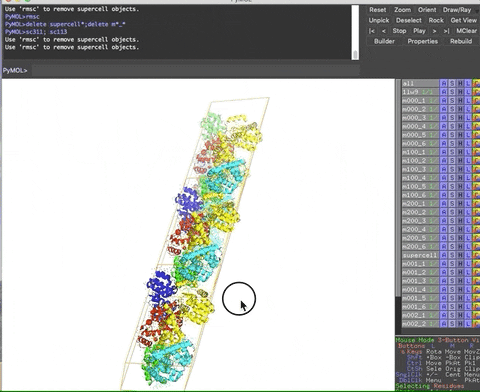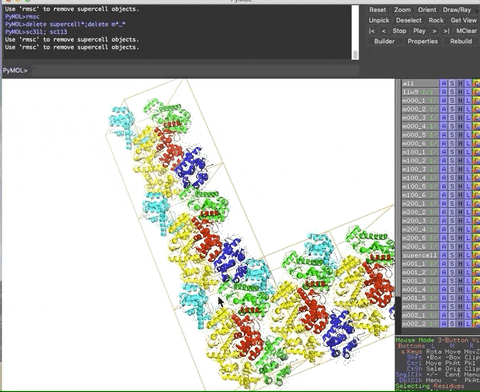
The variant file pymolschortuctsNobs4.py lacks a few functions that depend on the Python module beautifulsoup4 that is not shipped with PyMOL. Choose this variant file if you do not know how to add beautifulsoup4 to your version of PyMOL.
Selecting the right pymolshortcuts.py script
Have Open Source PyMOL or incentive version => 2.0.
If you want to use the shortcuts with a version of PyMOL that has a Python interpreter to which you can add an external module (generally any Open source PyMOL or the incentive version equal to or greater than 2.0), download the pymolshortcuts.py script.
Have incentive version < 2.0 or do not want to mess with installing an external module
If you have an incentive version less than 2.0 or you do not want to bother with the installation of the datetime module, download the pymolshortcutsNoDateTime.py. This version of the script lacks the functions that depend on the datetime module. By using this version You will not get error messages about this module not being found.
How to install the date time module
These instructions are only for the users of pymolshortcuts.py
The Python program pip is available for all installing external modules. The recommended way of using pip when more than one Python interpreter is on our computer is to import it as a module on the command line with the path of the python interpreter for which you are making the install. For example, to install the datetime module in the macports Python3.7, use sudo -H /opt/local/bin/python3.7 -m pip install --upgrade datetime. The minus m flag means import the following module. Depending on your configuration of macports, you may not need to use sudo -H.
Download the script file
Before you download the script file, be sure that you have set your system to show file extensions. Both Mac and Windows operating system hide the file extension by default.
On the Mac, do the following
- Open a new Finder window.
- In the menu bar, select Finder > Preferences.
- Check the box Show all filename extension
On Windows, do the following:
- Search for File Explorer Options in the Start search box and open the Folder Options box.
- Uncheck the box next to Hide extensions for known file types".
To download all of the files, git clone the repository.
Otherwise, left-click on one of the files above and left-click "Raw".
The raw file will be displayed in your browser.
Select save under the file pull-down of your browser.
Your browser may add .txt after .py.
Delete .txt by backspacing over it.
Then save the file to your home directory.
Your browser may try to append the .py with .txt again via a popup GUI, but do not select this option.

Installation
To have the shortcuts always available in PyMOL, add the command 'run ~/pymolshortcutsNobs4.py' (recommended for most users) or 'run ~/pymolshortcutsbs4.py' (for advanced users) to the .pymolrc, pymolrc or pymolrc.pml file in your home directory. This command will to load the functions in the script on every startup of PyMOL so you do not have to think about doing so again.
The pymolrc file is an optional file. You may have to create it with a text editor if you have not done so already. If you do not have a text editor, you can use PyMOL's built-in text editor. Go to File --> Edit pymolrc. The functions will be loaded into memory but will not be executed despite the use of the command run. You may want to store the script pymolshortcuts.py in a safer place than your home directory. I store mine in /Users/blaine/Scripts/PyMOLScripts/.
Configure the script for access to all shortcuts
Configuration: application start commands
A number of shortcuts open external programs. The commands that work on the Mac OS are active by default. These are not path-dependent. They should work regardless of the nonconventional location of your applications, as long as they are in your PATH. They are found around line 500 in the pymolshortcuts.py script file.
The commands for Mac OS need to be commented out and the commands for Linux or Windows need to be uncommented. Do not use a word processor to edit this file. Visual Studio Code is an excellent, free, and intuitive text editor that is platform independent. Install one of the Python in-plugins via the marketplace to get syntax highlighting of the pymolshortcuts.py script file
Configuration: paths to local directories and URLs of some web pages
Around line 500, there are also paths to local directories of PDB files and webmail sites that need to be customized.
Do not have PyMOL or do not want to mess with your existing PyMOL?
Note that multiple versions of PyMOL can be on one computer. On the Mac, you can give the package in Applications a different name. I append the version number to the stem of the file name, e.g., PyMOL233.app for version 2.3.3. Avoid trouble down the road by NOT introducing whitespaces into the file name.
Download the incentive version of the current PyMOL
The incentive version of PyMOL is the easiest to install due to the availability of installers. You can use the incentive version for 30 days for free without buying an annual license.
Open-source alternatives
Mac
See the instructions on the PyMOL Wiki page for installing on the Mac.
There are many ways of installing PyMOL on the Mac. I suggest using the install command below instead for installing pymol on the Mac with Macports. I have used Fink, Anaconda, Homebrew, and Macports. I have the fewest problems with Macports, followed by Fink.
The main barrier to using Macports is the need to install the Xcode-related command-line tools. The protocol for doing so varies with the version of the Mac operating system and the version of Xcode installed on your computer. Expect to spend at least an hour installing the command line tools and then Macports. If this sounds like too much trouble, download the 30-day trial of the incentive version.
Open source version of pymol via Macports works fine. It is missing a few minor thrills found in the incentive version like the ability to import background images into the viewport.
You can have your favorite protein rocking over a scene from your last vacation.
This is cool not essential for serious work.
Note that the installation of PyMOL via MacPorts on the Mac (sudo port install pymol +python37) allows specification of the version of the Python interpreter.
PyMOL can be installed with versions 2.7, 3.5, 3.6, and 3.7 of Python.
This is handy if you have a module written for only Python3.5 that you want to import into PyMOL to do something unique and creative without writing a new plugin for PyMOL.
However,


