Mzkit
Data toolkits for processing NMR, MALDI MSI, MALDI single cell, Raman Spectroscopy, LC-MS and GC-MS raw data, chemoinformatics data analysis and data visualization.
Install / Use
/learn @xieguigang/MzkitREADME
<span style="font-size: 3em;">Mzkit</span> 
Data toolkits for processing NMR, MALDI MSI, MALDI single cell, Raman Spectroscopy, LC-MS and GC-MS raw data, chemoinformatics data analysis and data visualization.
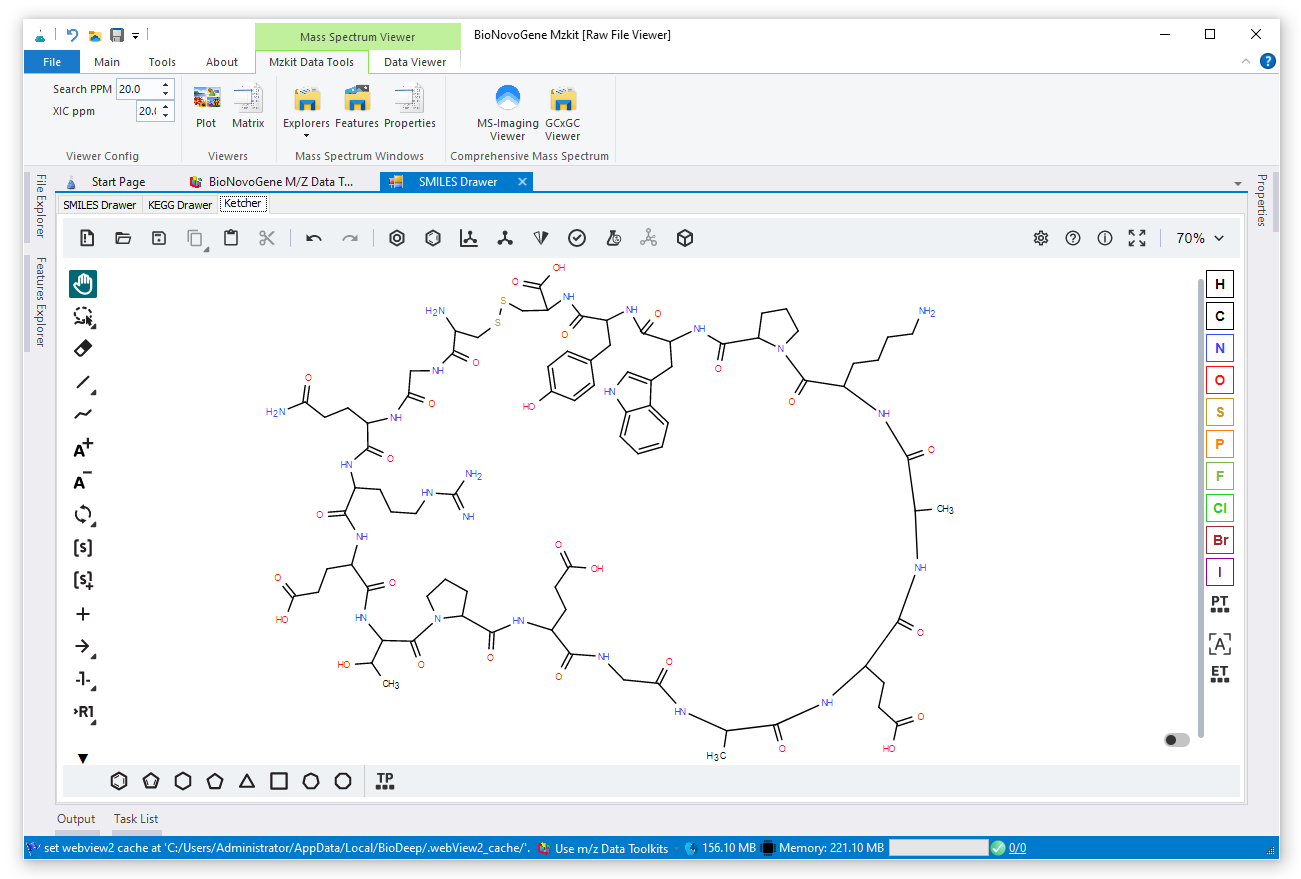
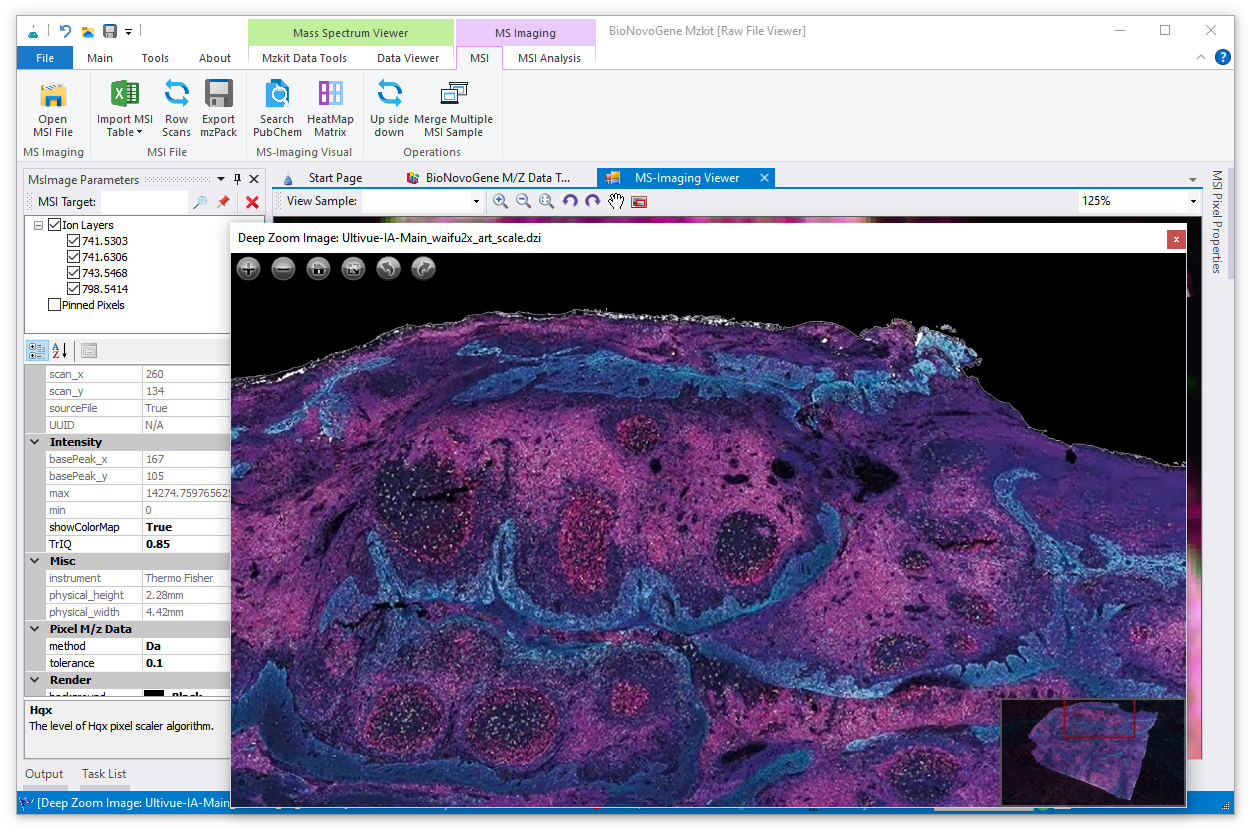
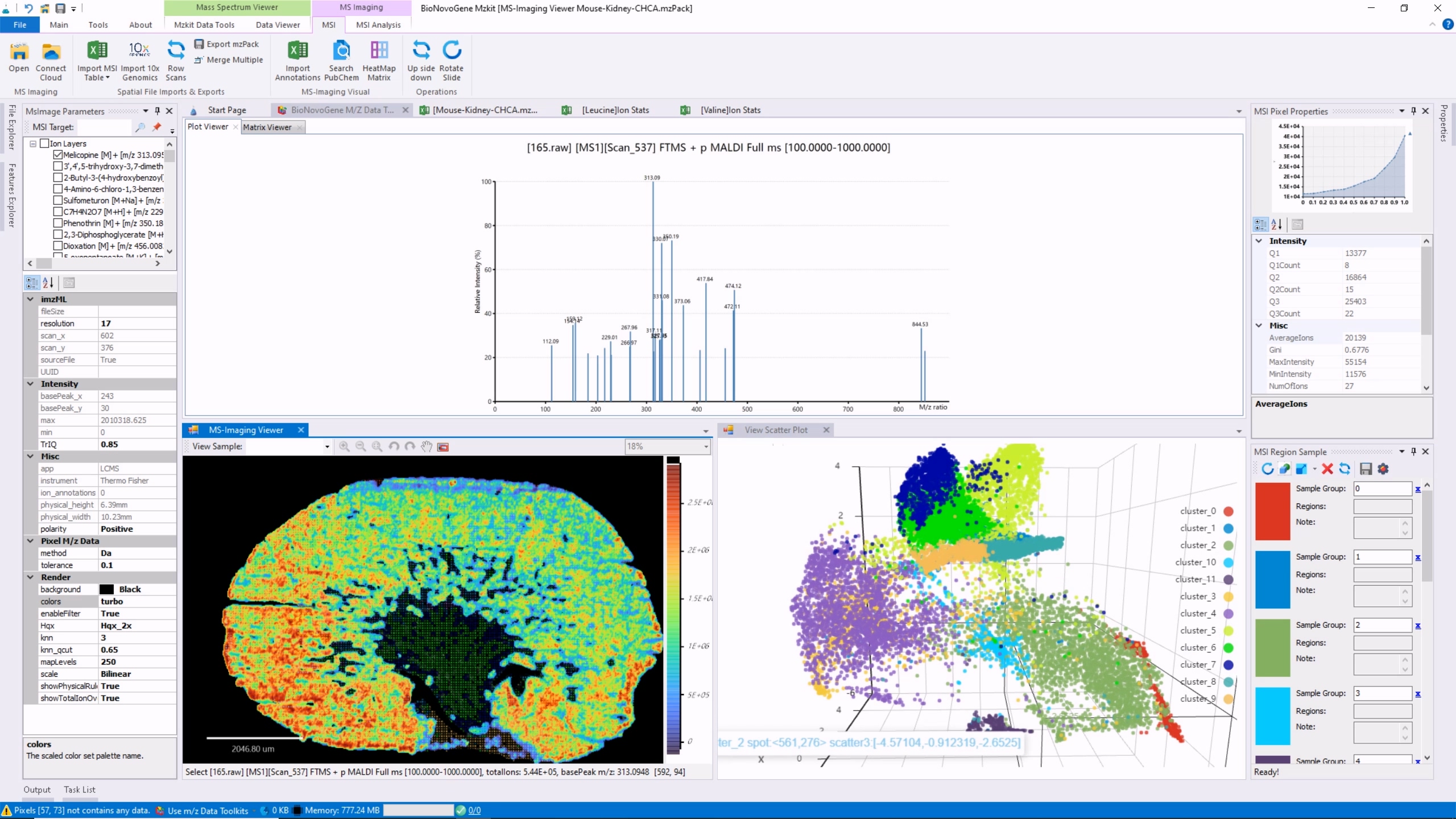
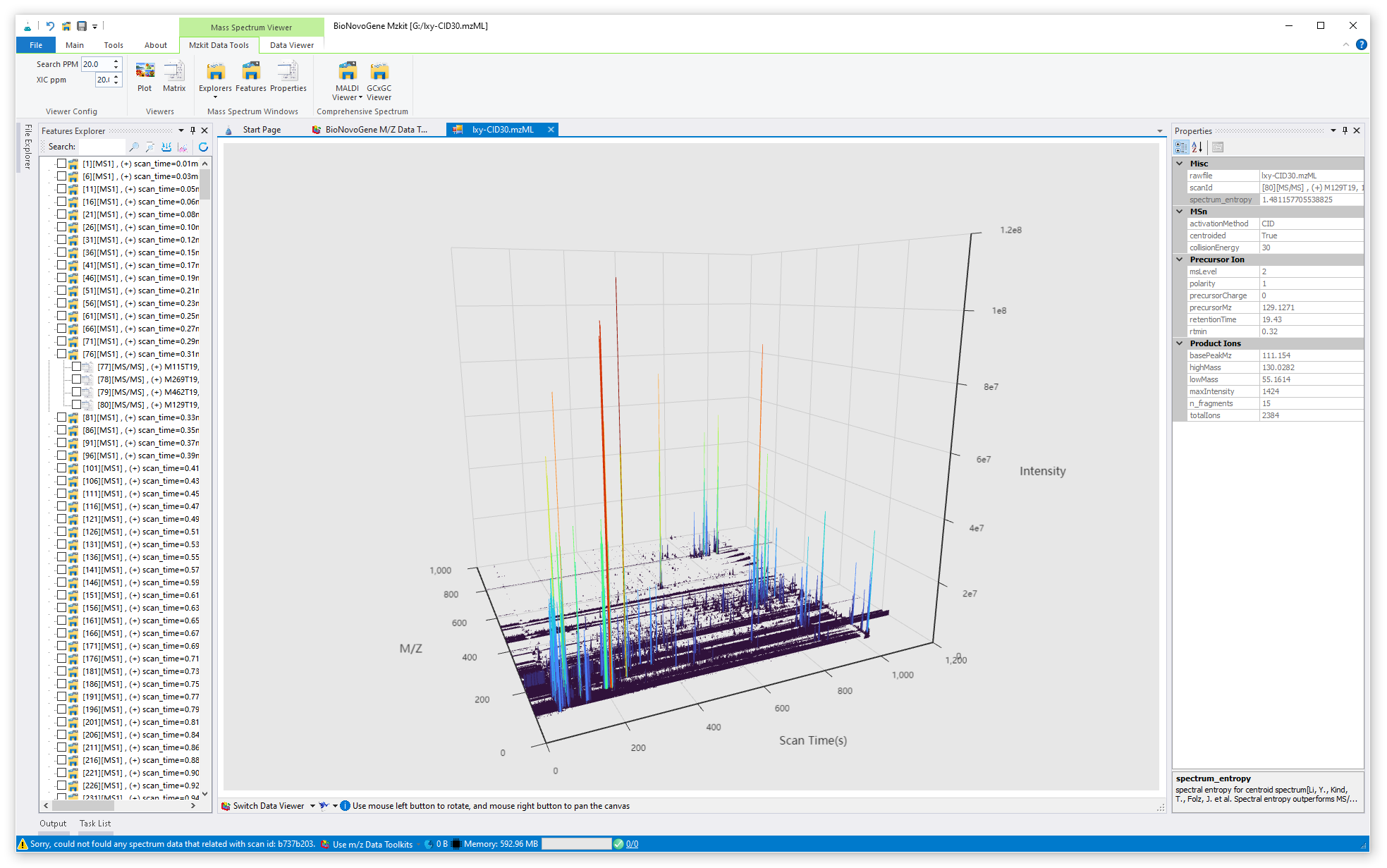
Mzkit is an open source raw data file toolkit for mass spectrometry data analysis, provides by the BioNovoGene corporation. The features of mzkit inlcudes: raw data file content viewer(XIC/TIC/Mass spectrum plot/MS-Imaging), build molecule network, formula de-novo search, de-novo annotation of the unknown metabolite features and targeted data quantification.
- View User Manual PowerPoint
- View Mzkit Update History
- Found me at: https://bio.tools/mzkit
Download Software: https://mzkit.org/download.html
Docker Image: docker pull xieguigang/mzkit
LICENSE
- MZKit® is a registered trademark of BioNovoGene Corporation, protected by copyright law and international treaties.
- RawFileReader reading tool. Copyright © 2016 by Thermo Fisher Scientific, Inc. All rights reserved.
- MS-DIAL was launched as a universal program for untargeted metabolomics that supports multiple instruments (GC/MS, GC/MS/MS, LC/MS, and LC/MS/MS) and MS vendors (Agilent, Bruker, LECO, Sciex, Shimadzu, Thermo, and Waters). Part of the spectrum algorithm in MZKit is developed based on the MS-DIAL project.
MIT License
Copyright (c) 2025 xieguigang@metabolomics.ac.cn / gg.xie@bionovogene.com, BioNovoGene Co., LTD.
Permission is hereby granted, free of charge, to any person obtaining a copy
of this software and associated documentation files (the "Software"), to deal
in the Software without restriction, including without limitation the rights
to use, copy, modify, merge, publish, distribute, sublicense, and/or sell
copies of the Software, and to permit persons to whom the Software is
furnished to do so, subject to the following conditions:
The above copyright notice and this permission notice shall be included in all
copies or substantial portions of the Software.
THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE
AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM,
OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE
SOFTWARE.
-
-
- 1.1.1. Background Task
-
1.2. Search Feature
-
1.3. XIC plot
- 1.3.1. XIC overlay
-
1.4. TIC plot
-
1.5. Raw Scatter
-
1.6. View Mass spectra
-
1.7. Save Plot and Export matrix
- 1.7.1. Export XIC
-
1.8. Ms-Imaging
-
-
-
2.1. Formula search
- 2.1.1. Export Formula Search Result
-
2.2. Molecular Networking
- 2.2.1. step1 select ions
- 2.2.2. step2 build network
- 2.2.3. step3 view network data
- 2.2.4. step4 network visualization
- 2.2.5. Export Network Data
- 2.2.6. Save Network Visual
-
-
- 3.1. Switch Between Toolkit
- 3.2. Install Mzkit
- 3.3. Uninstall Mzkit
Credits
This open source mass spectrometry data toolkit is developed at the BioDeep R&D laboratory and brought to you by
BioNovoGenecorporation.






1. <a name='RawDataViewerInstruction'></a>Raw Data Viewer Instruction
Important Note: this application only supports the open source mzXML/mzML/imzML raw data file formats. For view the data in the vendor format file like the Thermo *.Raw required convert to the mzXML file format at first. It is recommended that convert the vendor format file to mzXML via ProteoWizard.
1.1. <a name='Importsrawdatafile'></a>Imports raw data file
For view the file content of the mzXML or mzML datafile in mzkit, you must imports the raw data file into the mzkit at first. Here is how: select the Main tabpage of mzkit you will see the Open command for the raw data imports operation. Then you are going to click this Open command button, choose the raw data file for imports and wait for it finished.

Then you should see the files that you've imports into mzkit on the File Explorer dock panel if there is no error occurs during the raw data file imports progress. Now you can click on the raw data file tree to expend it and click one of the feature in your raw file to view the content data.

1.1.1. <a name='BackgroundTask'></a>Background Task
For improvements of the user experience when you are using the mzkit application, the raw data files is imports to mzkit application under a background task. You can see the background task progress throught the Task List window:
1.2. <a name='SearchFeature'></a>Search Feature
the search bar on the top of the file tree is the m/z search input: you can input a specific m/z value or formula expression in the search bar for search the matched features in your raw data file. This operation is usually apply for the XIC data search.
When you have click on the search button, then all of the M/z feature in your raw data file that match the ppm condition will be listed in the Featrue List dock panel:

An example of search m/z 834.6 with tolerance error 30ppm. the result in the
Feature Listis usually used for create a XCI plot.
1.3. <a name='XICplot'></a>XIC plot
Expends the file content tree in the File Explorer, and then mouse right click on one MS2 feature in your file, select XIC for create a XIC plot for a specific ion feature:

<div style="page-break-after:always;"></div>The XIC plot is a kind of time-signal chromatography plot of a specific m/z ion.
1.3.1. <a name='XICoverlay'></a>XIC overlay
you can click on the checkbox besides the Ms2 feature for select different ion feature for create the XIC overlay plot:

1.4. <a name='TICplot'></a>TIC plot
as the same as create a XIC plot, you also can create TIC plot for a single file or multiple file by select multiple file by check on checkbox:

<div style="page-break-after:always;"></div>The TIC plot is similar to the XIC plot, data is generated from all ions.
1.5. <a name='RawScatter'></a>Raw Scatter
just click on the node of the raw file, then you will open the raw scatter plot of your specific raw data file, example as:
1.6. <a name='ViewMassspectra'></a>View Mass spectra
For view the mass spectra data in your file, just click on one of the scan feature in your raw data file:

1.7. <a name='SavePlotandExportmatrix'></a>Save Plot and Export matrix
The mzkit application provides the function for save the plot image and the plot data in your raw data file. for example, select the Data Viewer tab page in mzkit, you will see two viewer action buttons in the menu:
- [
Snapshot] for export the XIC/TIC/MS2 data plot image to a specific file. - [
Save Matrix] for export the Mass spectra or Chromatography data to a specific Excel table file.

clawhub
331.2kUse the ClawHub CLI to search, install, update, and publish agent skills from clawhub.com
codebase-memory-mcp
843High-performance code intelligence MCP server. Indexes codebases into a persistent knowledge graph — average repo in milliseconds. 64 languages, sub-ms queries, 99% fewer tokens. Single static binary, zero dependencies.