Parallax
Tool for interactive embeddings visualization
Install / Use
/learn @uber-research/ParallaxREADME
Parallax
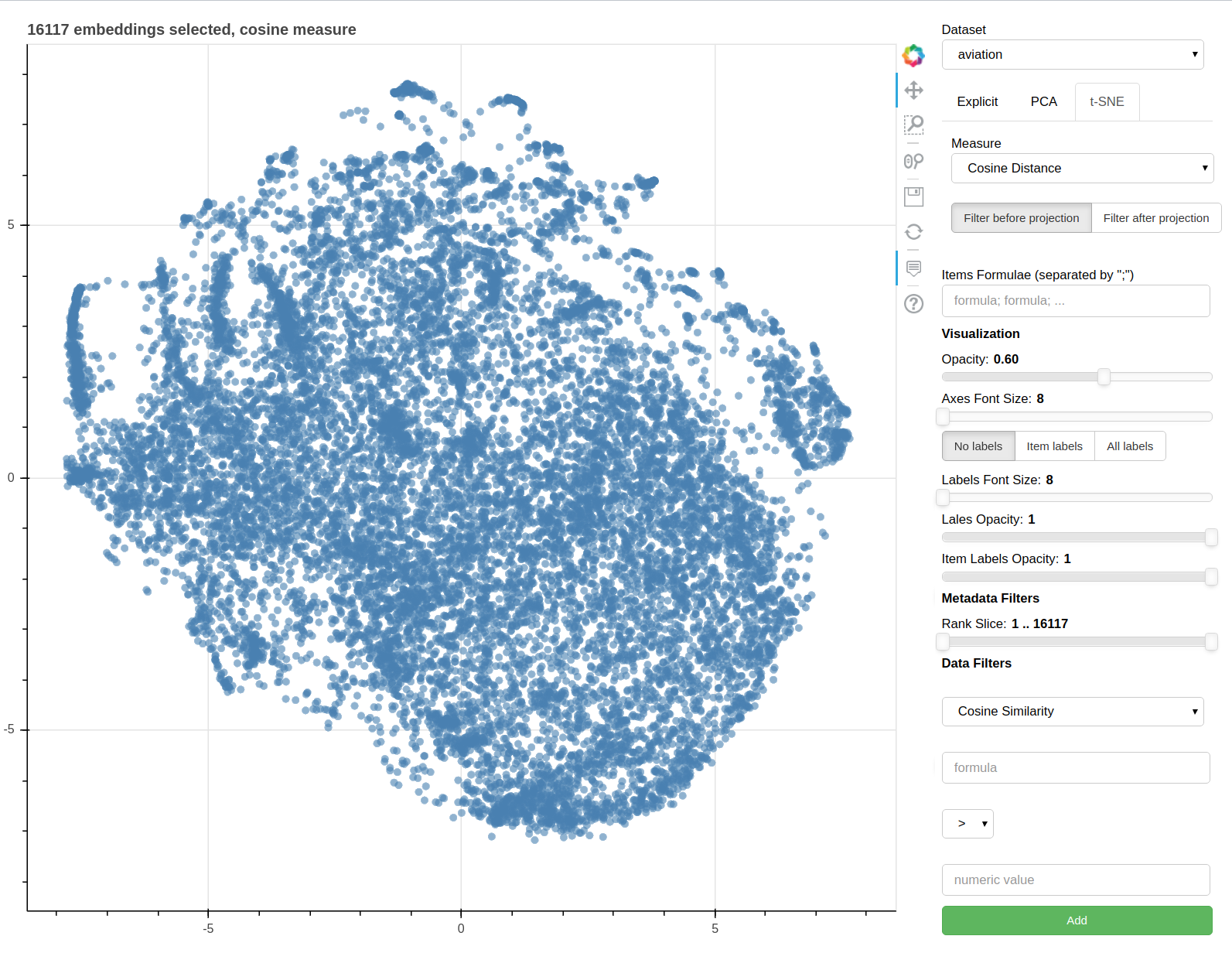
Parallax is a tool for visualizing embeddings.
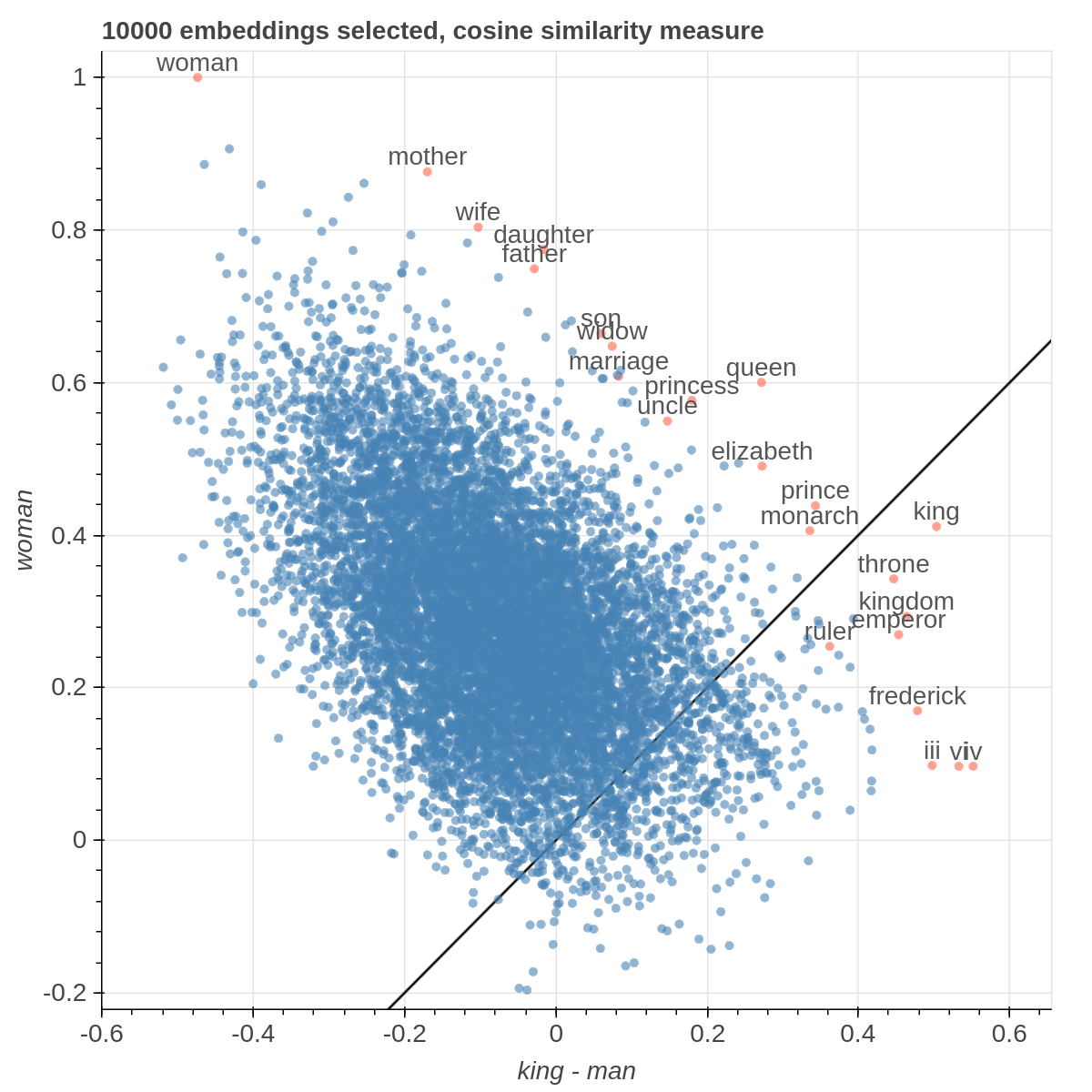
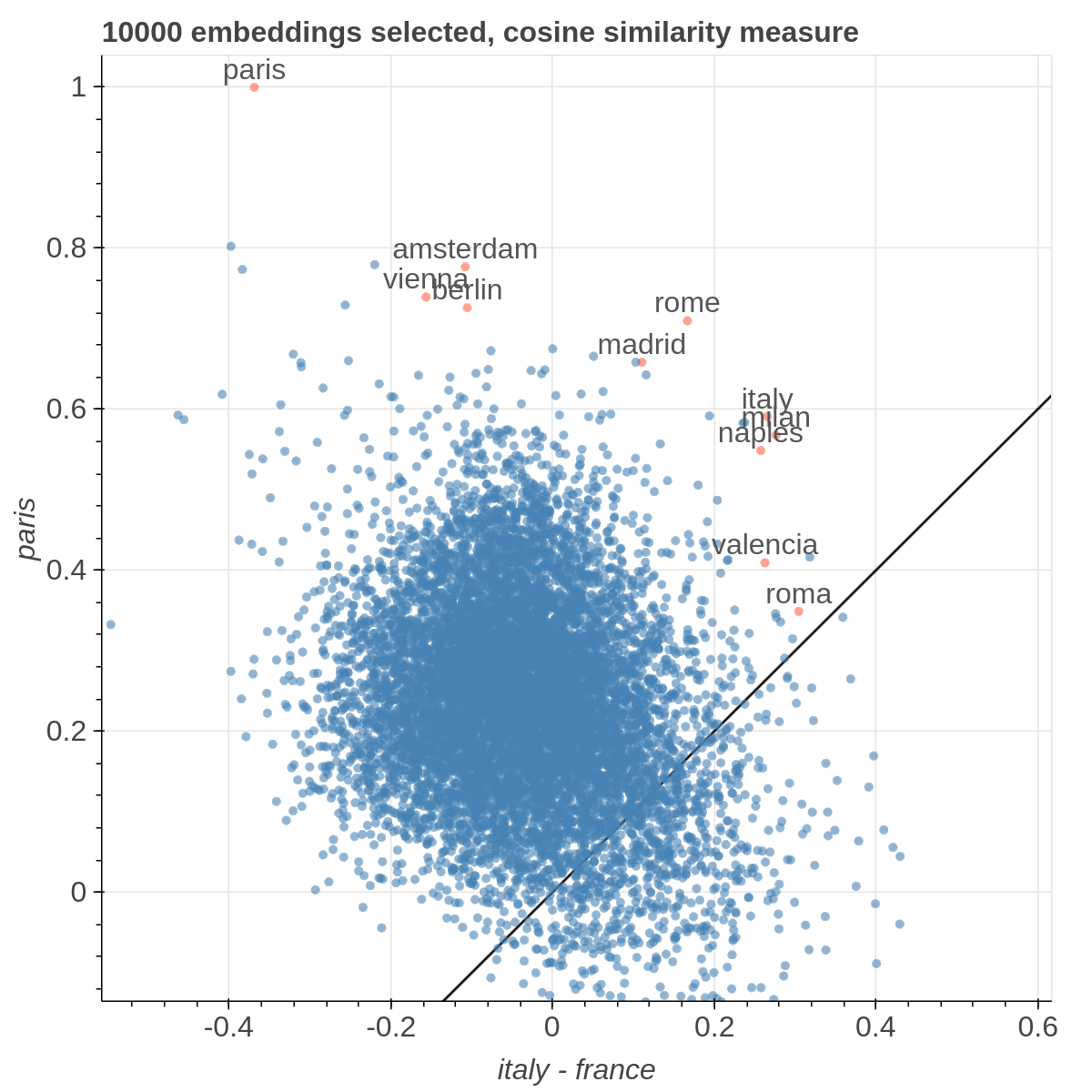
It allows you to visualize the embedding space selecting explicitly the axis through algebraic formulas on the embeddings (like king-man+woman) and highlight specific items in the embedding space.
It also supports implicit axes via PCA and t-SNE.
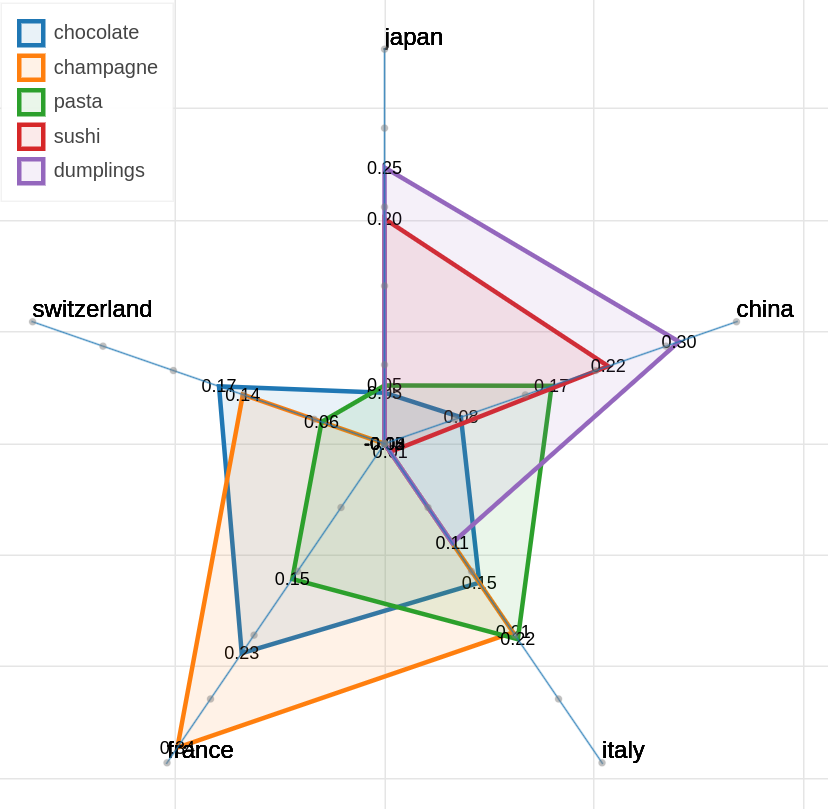
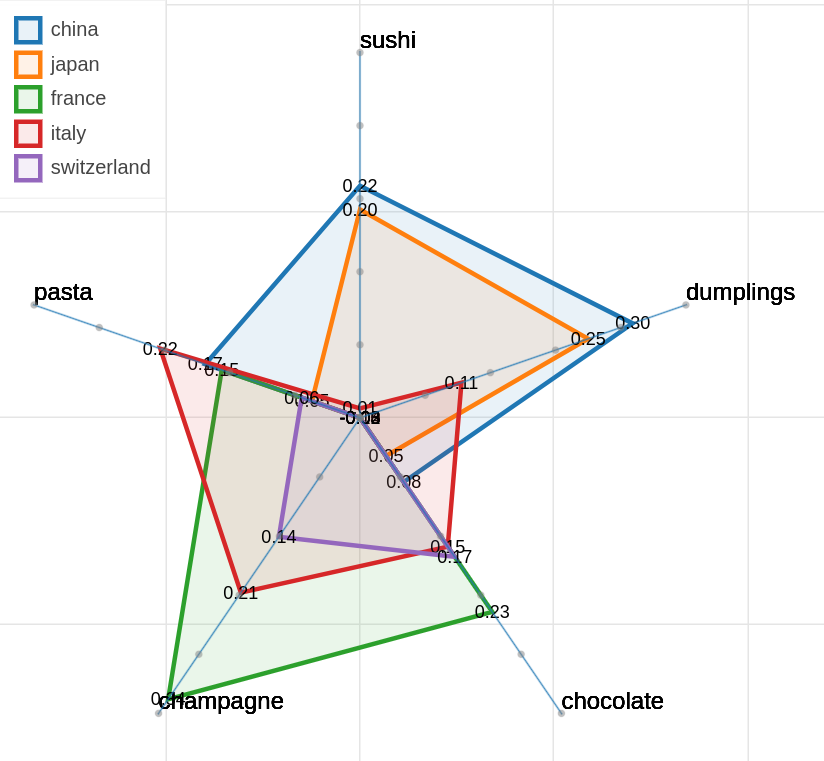
There are three main views: the cartesian view that enables comparison on two user defined dimensions of variability (defined through formulae on embeddings), the comparison view that is similar to the cartesian but plots points from two datasets at the same time, and the polar view, where the user can define multiple dimensions of variability and show how a certain number of items compare on those dimensions.
This repository contains the code used to obtain the visualization in: Piero Molino, Yang Wang, Jiwei Zhang. Parallax: Visualizing and Understanding the Semantics of Embedding Spaces via Algebraic Formulae. ACL 2019.
And extended version of the paper that describes thoroughly the motivation and capabilities of Parallax is available on arXiv
If you use the tool for you research, please use the following BibTex for citing Parallax:
@inproceedings{
author = {Piero Molino, Yang Wang, Jiwei Zhang},
booktitle = {ACL},
title = {Parallax: Visualizing and Understanding the Semantics of Embedding Spaces via Algebraic Formulae},
year = {2019},
}
The provided tool is a research prototype, do not expect the degree of polish of a final commercial product.
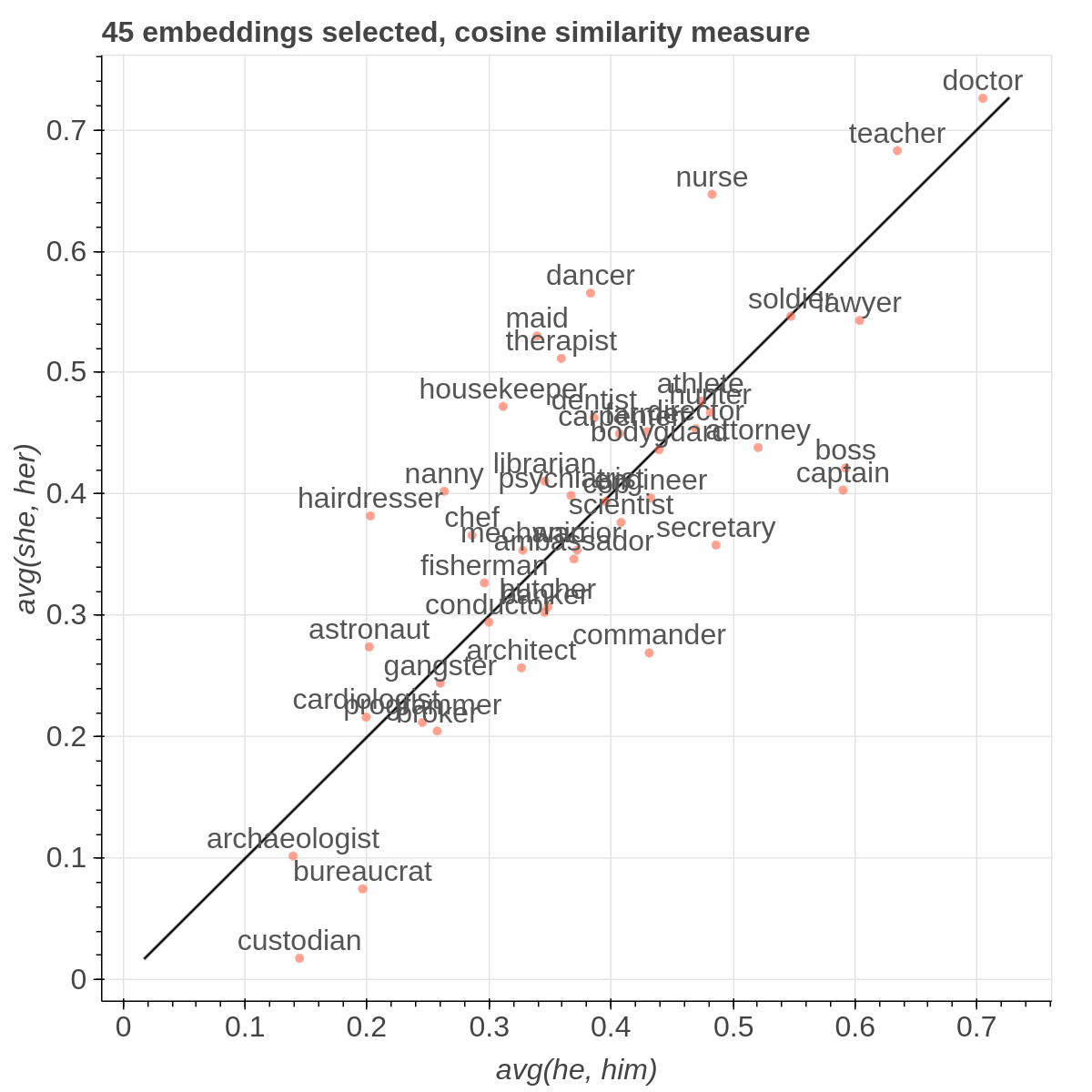
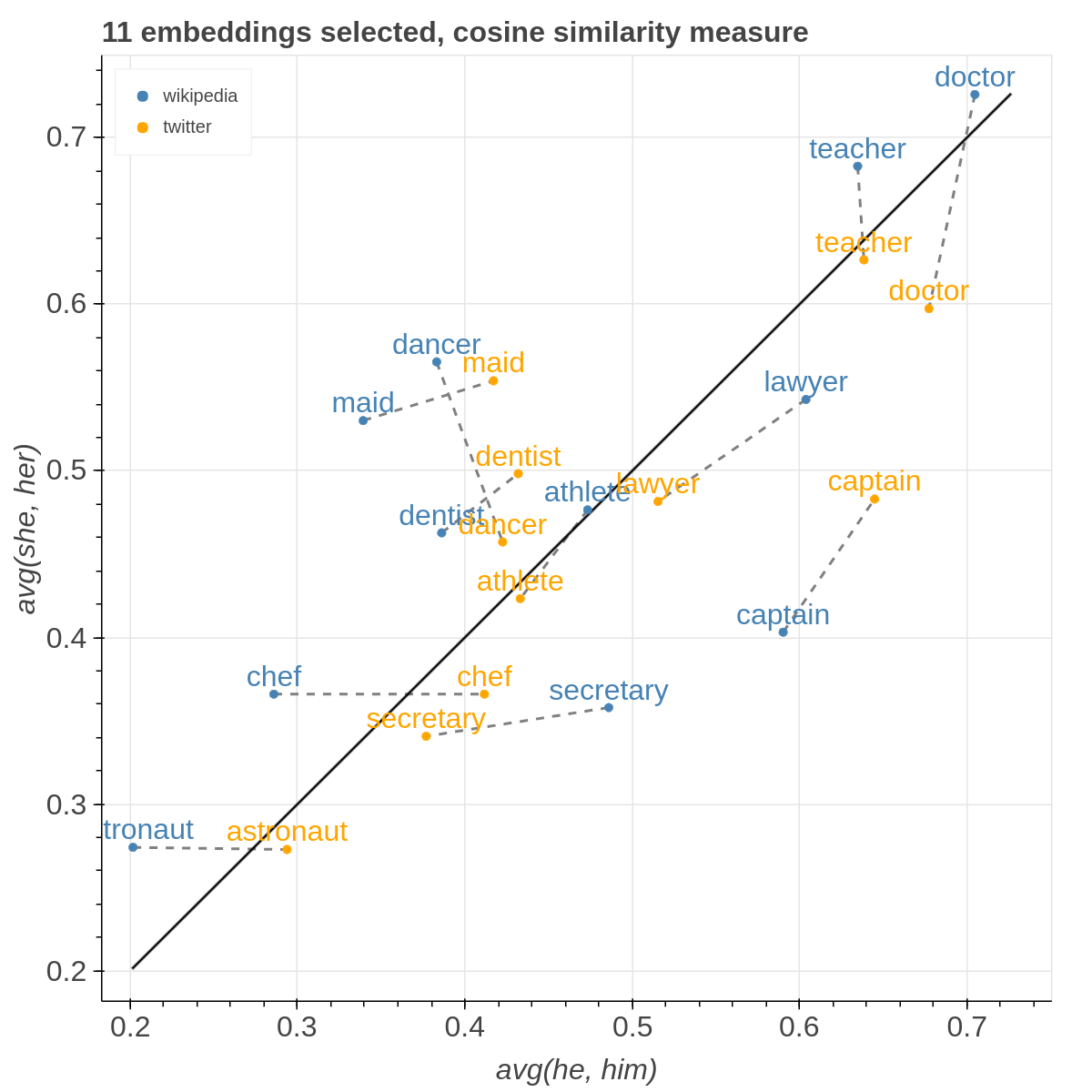
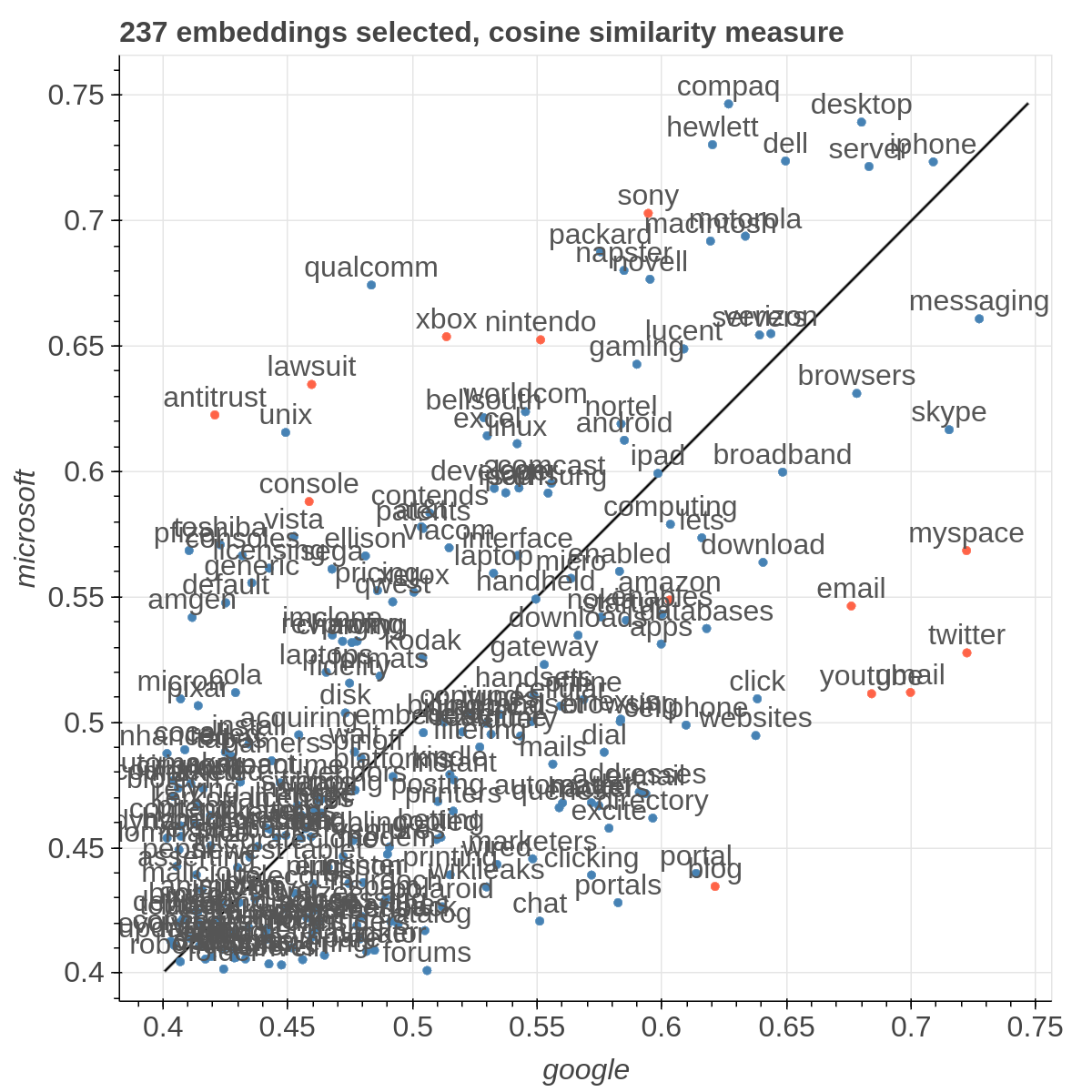
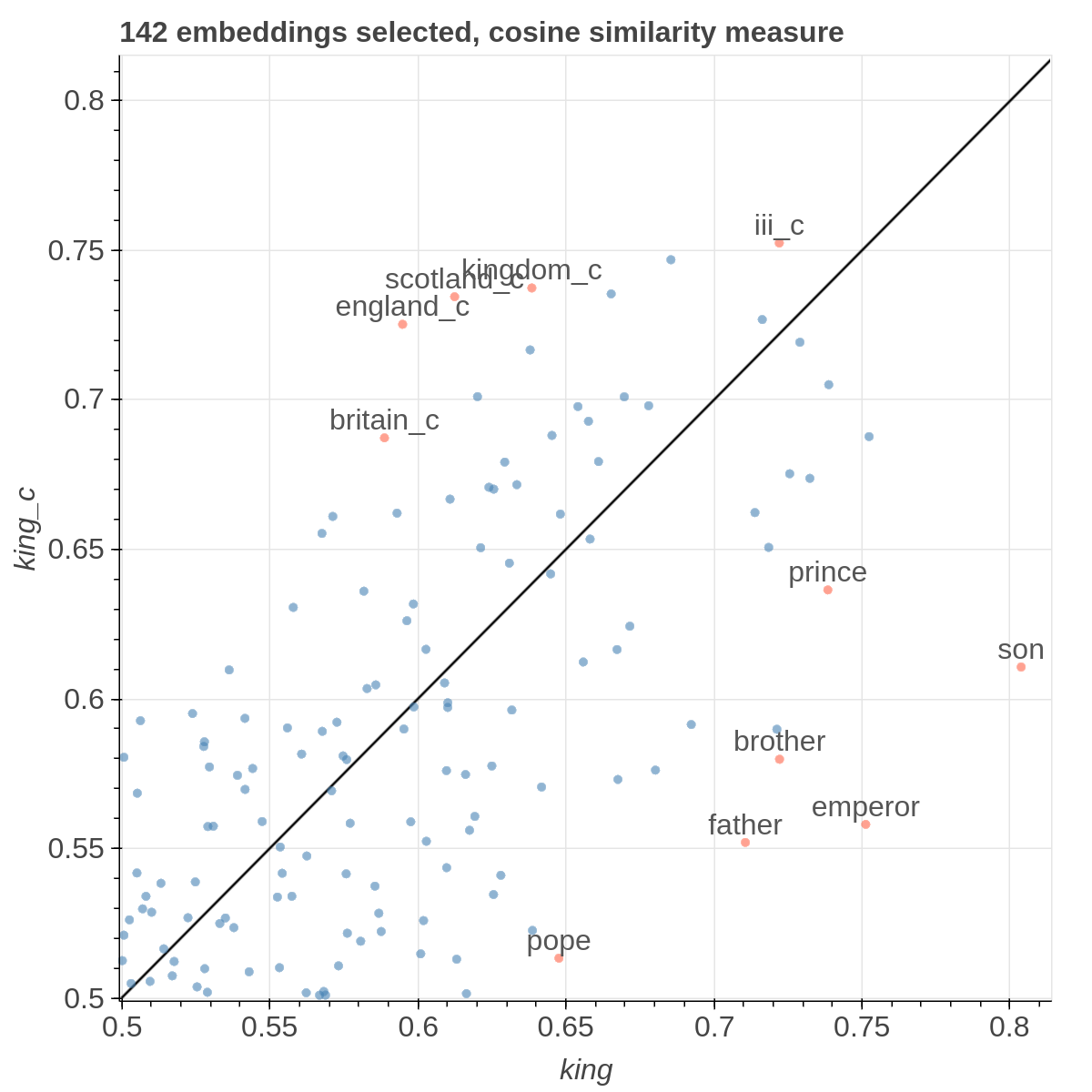
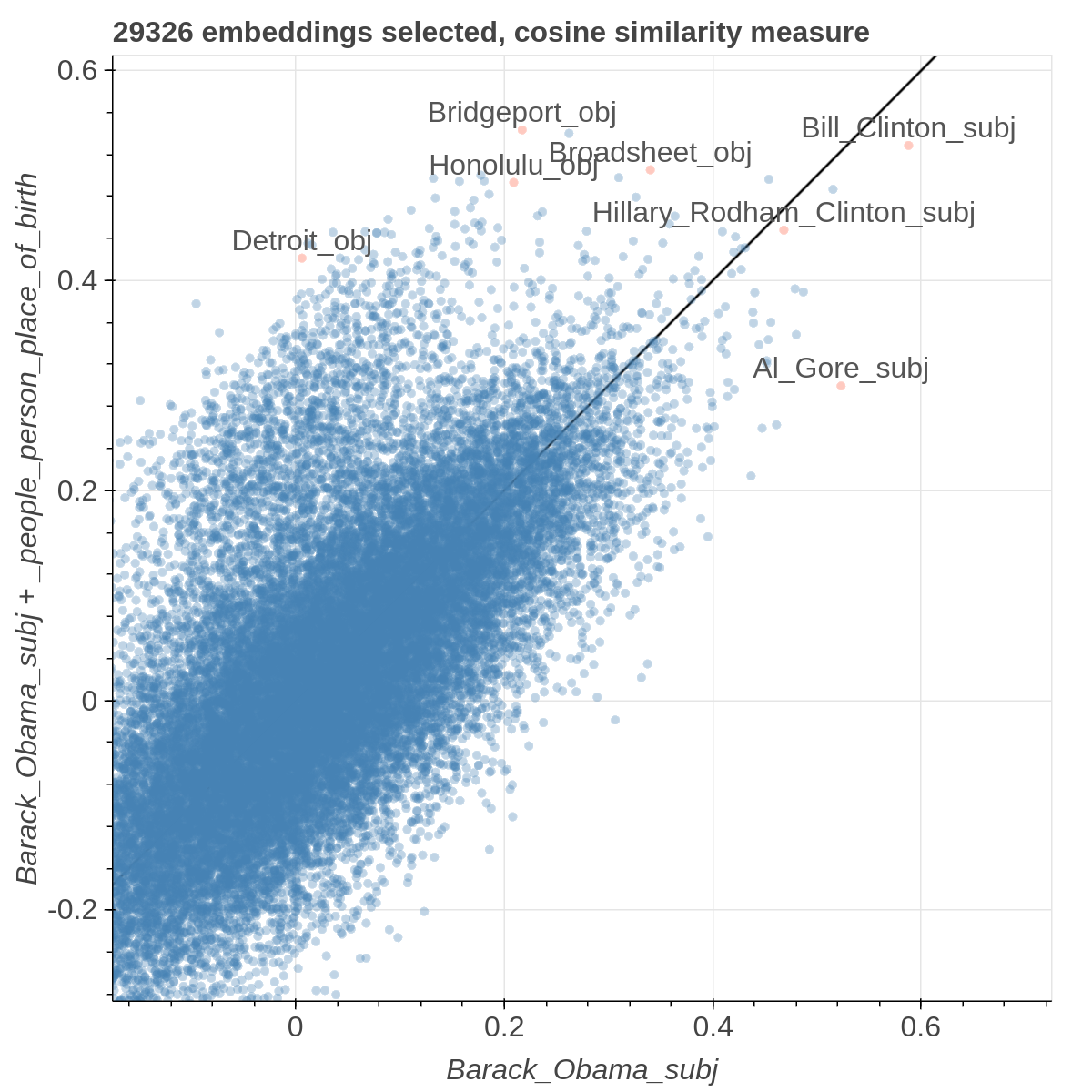
Example visualizations
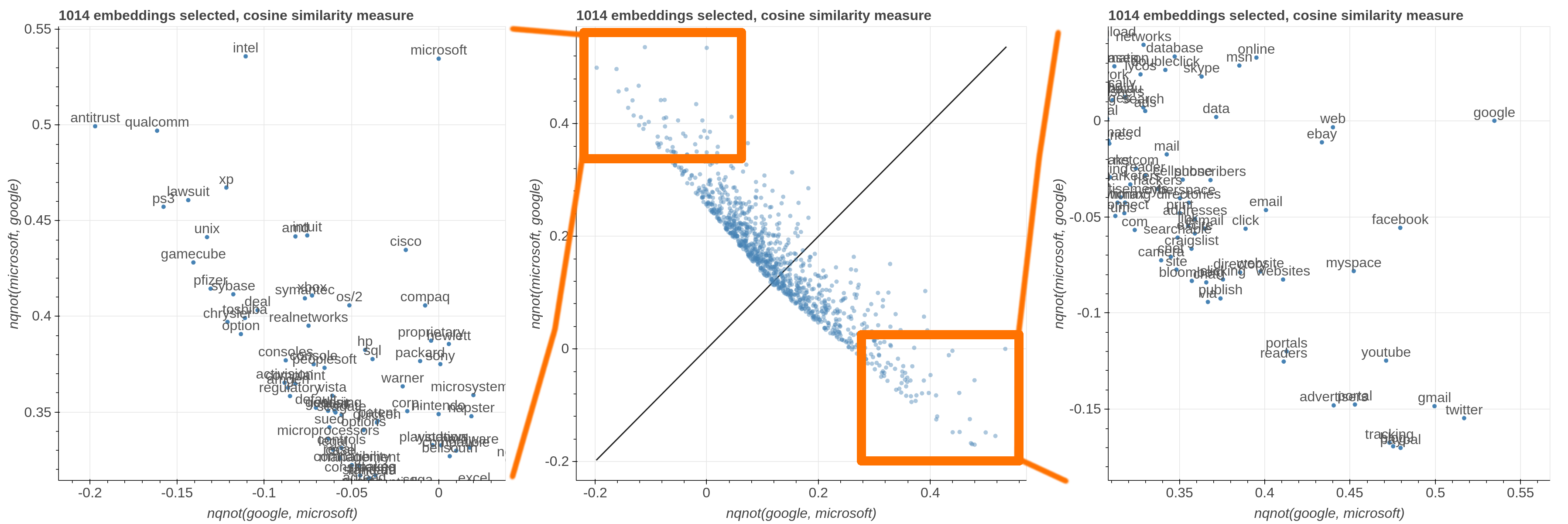
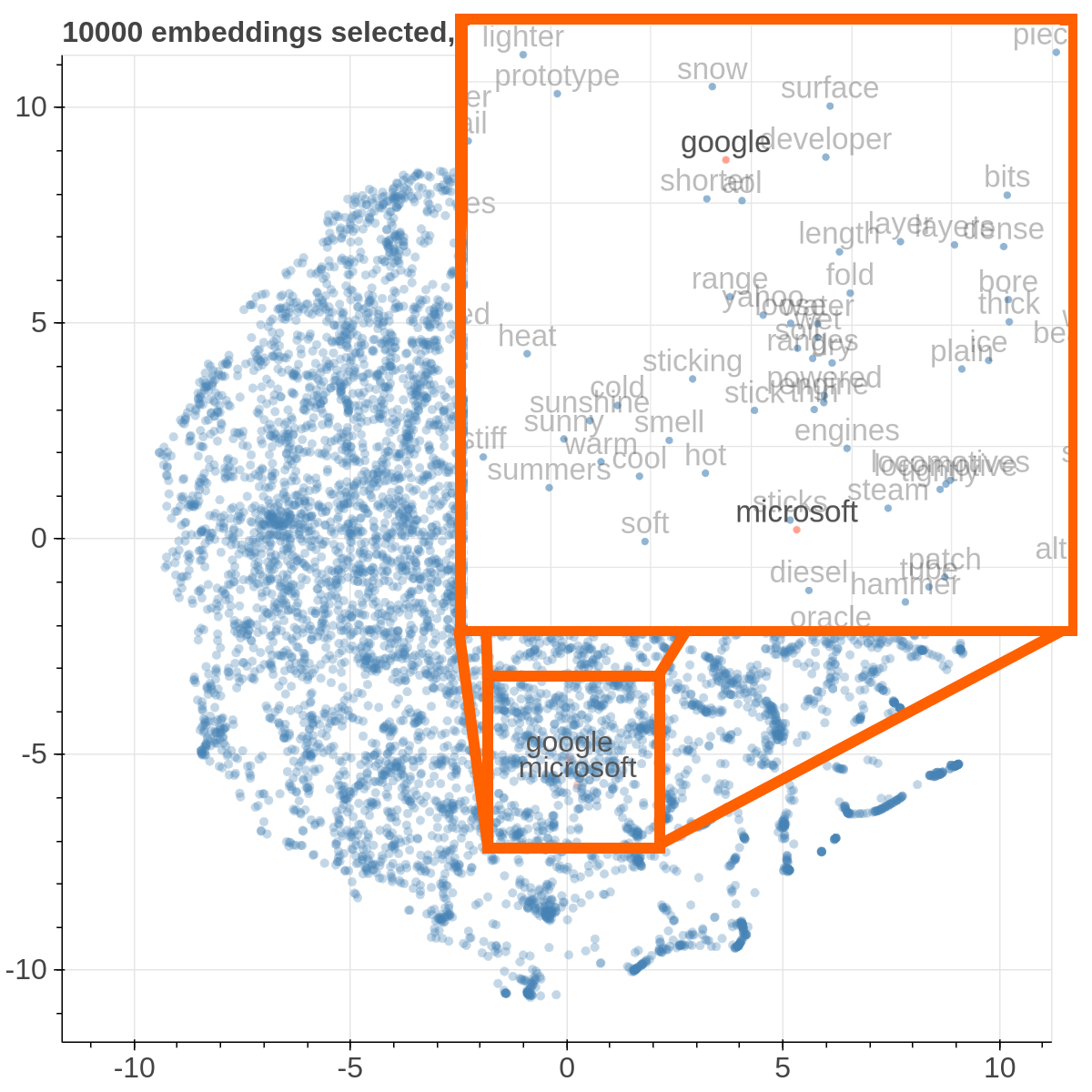
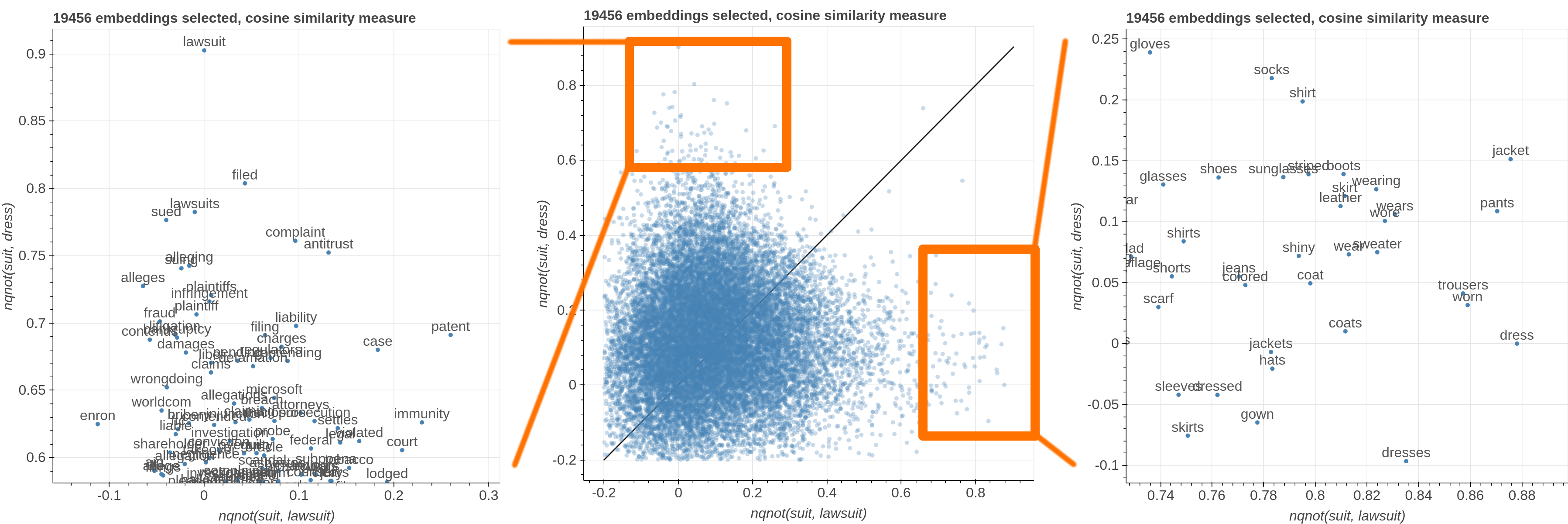
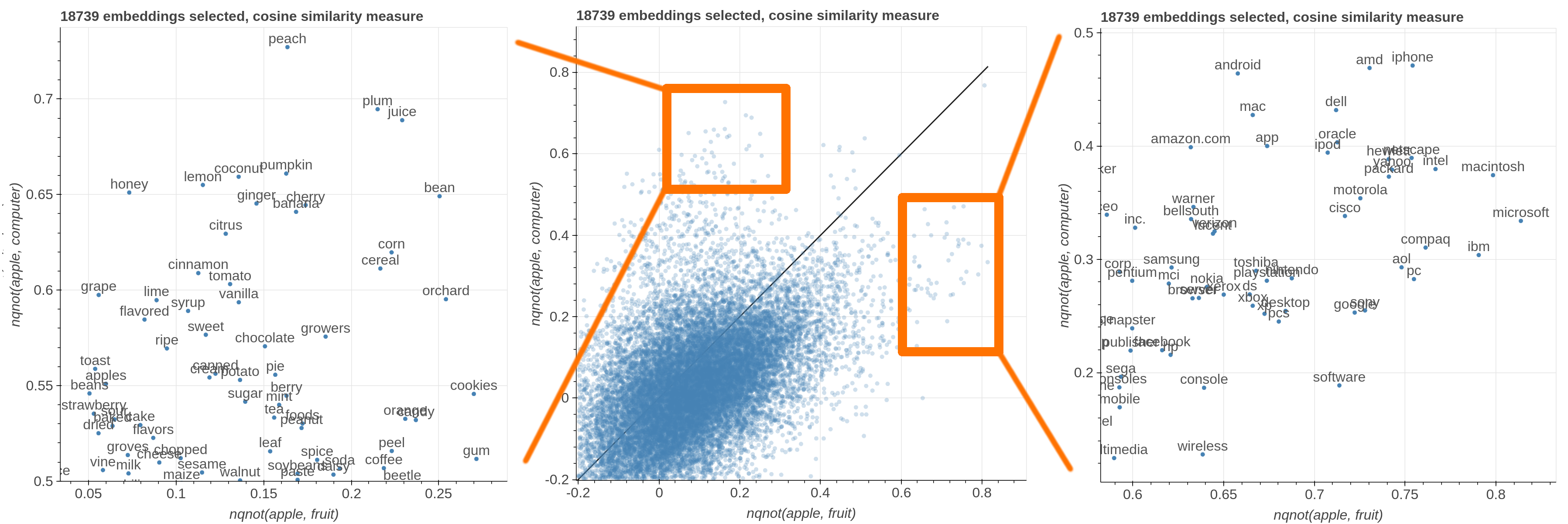
Here are some samples visualizations you can obtain with the tool. If you are interested in the details and motivation for those visualizations, please read the extended paper.
Set Up Environment (using virtualenv is not required)
virtualenv -p python3 venv
. venv/bin/activate
pip install -r requirements.txt
Download example data
In order to replicate the visualizations in our paper, you can download the GloVe embeddings for Wikipedia + Gigaword and twitter from the GloVe website. In the paper we used 50 dimensional embeddings, but feel free to experiment with embeddings with more dimensions. The two files you'll need specifically are:
- http://nlp.stanford.edu/data/glove.6B.zip
- http://nlp.stanford.edu/data/glove.twitter.27B.zip
After you unzipped them, create a data folder and copy glove.6B.50d.txt and glove.twitter.6B.50d.txt inside it.
In order to obtain the metadata (useful for filtering by part of speech for instance), use the automatic script in modules/generate_metadata.py. If you are uncertain about how to use it, the input format is the same of the main scripts as described afterwards, you can learn about the parameters by running python modules/generate_metadata.py -h.
If you placed the correct files in the data directory, after you run generate_metadata you'll find two additional JSON files in the data directory containing metadata for both sets of embeddings.
Run
To obtain the cartesian view run:
bokeh serve --show cartesian.py
To obtain the comparison view run:
bokeh serve --show comparison.py
To obtain the polar view run:
bokeh serve --show polar.py
You can add additional arguments like this:
bokeh serve --show cartesian.py --args -k 20000 -l -d '...'
-dor--datasetsloads custom embeddings. It accepts a JSON string containing a list of dictionaries. Each dictionary should contain a name field, an embedding_file field and a metadata_file field. For example:[{"name": "wikipedia", "embedding_file": "...", "metadata_file": "..."}, {"name": "twitter", "embedding_file": "...", "metadata_file": "..."}].nameis just a mnemonic identifier that is assigned to the dataset so that you can select it from the interface,embedding_fileis the path to the file containing the embeddings,metadata_fileis the path that contains additional information to filter out the visualization. As it is a JSON string passed as a parameter, do not forget to escape the double quotes:
bokeh serve --show cartesian.py --args "[{\"name\": \"wikipedia\", \"embedding_file\": \"...\", \"metadata_file\": \"...\"}, {\"name\": \"twitter\", \"embedding_file\": \"...\", \"metadata_file\": \"...\"}]""
-kor--first_kloads only the firstkembeddings from the embeddings files. This assumes that the embedding in those files are sorted by unigram frequency in the dataset used for learning the embeddings (that is true for the pretrained GloVe embeddings for instance) so you are loading the k most frequent ones.-lor--lablesgives you the option to show the level of the embedding in the scatterplot rather than relying on the mousehover. Because of the way bokeh renders those labels, this makes the scatterplot much slower, so I suggest to use it with no more than 10000 embeddings. The comparison view requires at least two datasets to load.
Custom Datasets
If you want to use your own data, the format of the embedding file should be like the GloVe one:
label1 value1_1 value1_2 ... value1_n
label2 value2_1 value2_2 ... value2_n
...
while the metadata file is a json file that looks like the following:
{
"types": {
"length": "numerical",
"pos tag": "set",
"stopword": "boolean"
},
"values": {
"overtones": {"length": 9, "pos tag": ["Noun"], "stopword": false},
"grizzly": {"length": 7, "pos tag": ["Adjective Sat", "Noun"], "stopword": false},
...
}
}
You can define your own type names, the supported data types are boolean, numerical, categorical and set.
Each key in the values dictionary is one label in the embeddings file and the associated dict has one key for each type name in the types dictionary and the actual value for that specific label.
More in general, this is the format of the metadata file:
{
"types": {
"type_name_1": ["numerical" | "binary" | categorical" | "set"],
"type_name_2": ["numerical" | "binary" | categorical" | "set"],
...
},
"values": {
"label_1": {"type_name_1": value, "type_name_2": value, ...},
"label_2": {"type_name_1": value, "type_name_2": value, ...},
...
}
}
User Interface
Cartesian View
The side panel of the cartesian view contains several controls.
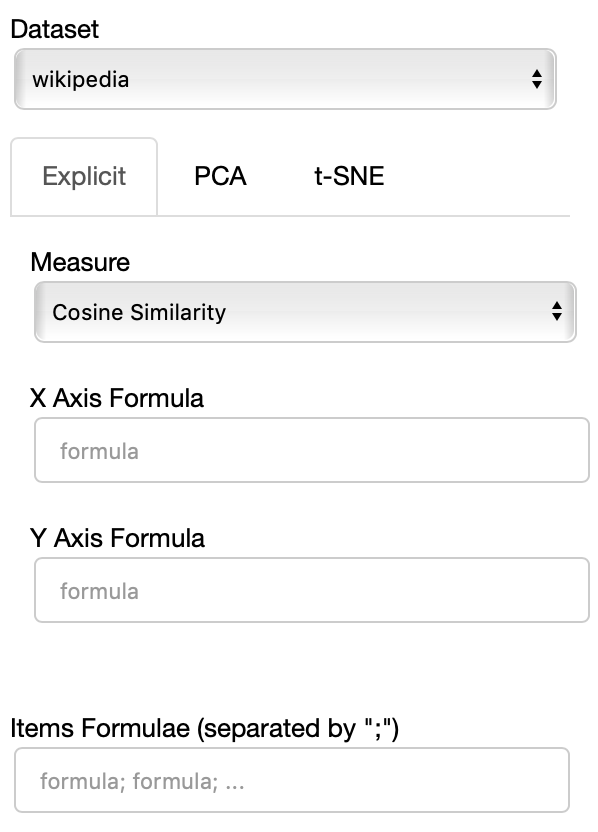
The Dataset dropdown menu allows you to select which set of embeddings use use among the ones loaded.
The names that will appear in the dropdown menu are the names specified in the JSON provided through the --datasets flag.
If the Explicit projection method is selected, the user can specify formulae as axes of projection (Axis 1 and Axis 2 fields.
Those formulae have embeddings labels as atoms and can contain any mathematical operator interpretable by python. Additional operators provided are
avg(word[, word, ...])for computing the average of a list of embeddingsnqnot(words, word_to_negate)which implements the quantum negation operator described in Dominic Widdows, Orthogonal Negation in Vector Spaces for Modelling Word-Meanings and Document Retrieval, ACL 2003
The Measure field defines the measure to use to compare all the embedding to the axes formulae.
Moreover, the user can select a subset of items to highlight with a red dot instead of a blue one. They will also have their dedicated visualization controls to make them more evident. Those items are provided in the Items field, separated by a semicolon. The items can be formulae as described above, which includes also single words.
If the PCA projection method is selected, the user can select how the filters (explained later) are applied, if before or after the projection.
