Whippet.jl
Lightweight and Fast; RNA-seq quantification at the event-level
Install / Use
/learn @timbitz/Whippet.jlREADME
Whippet
Graphical Overview
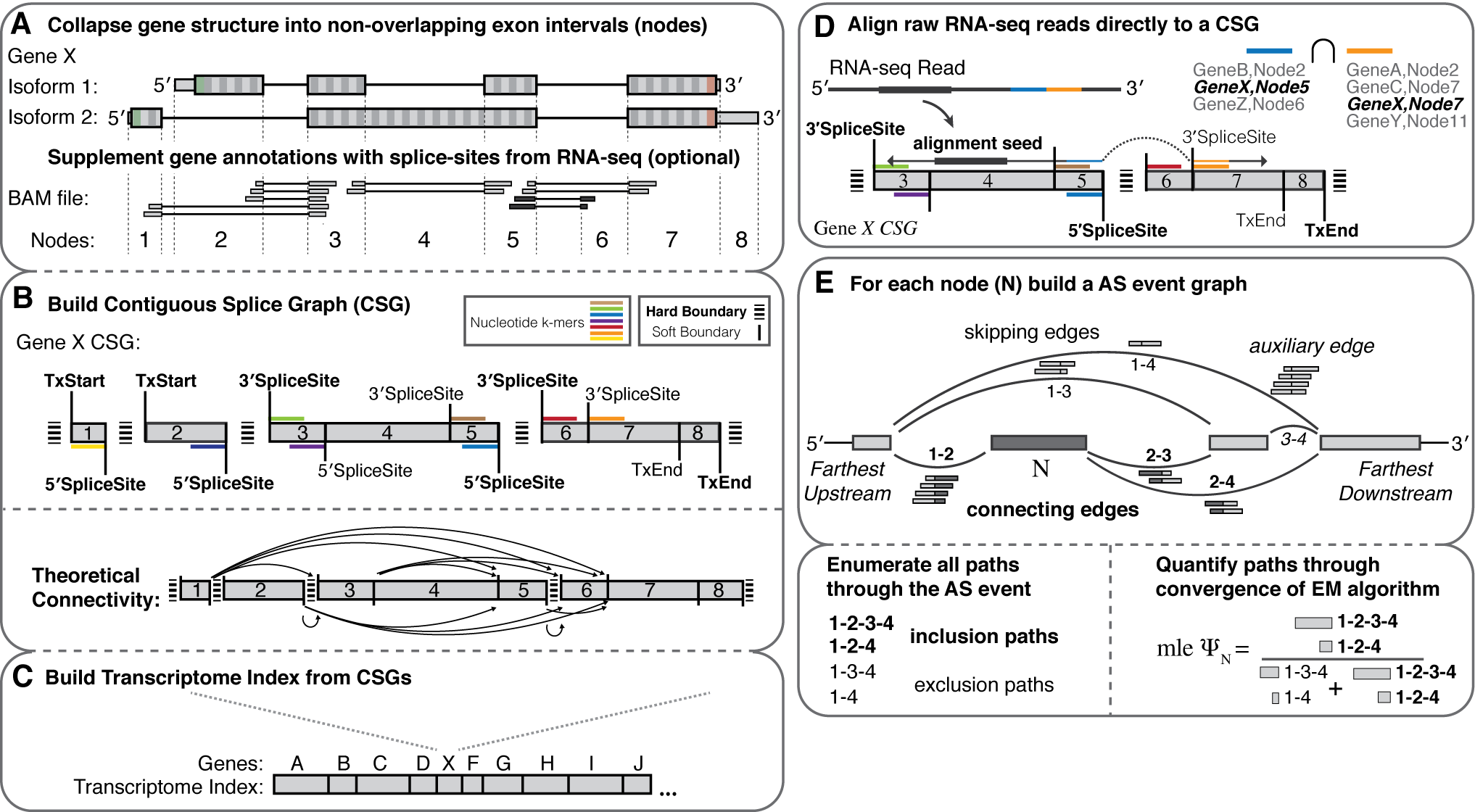
Features
- Splice graph representations of transcriptome structure
- Build an index for any species with a genome and annotation file
- Supplement the index with splice-sites/exons from independently aligned RNA-seq (BAM file).
- de novo AS event discovery (between indexed donor/acceptor splice sites)
- High speed PolyA+ Spliced Read Alignment (Read lengths <= 255)
- Repetitive read assignment for gene families
- Bias correction methods for 5' sequence and GC-content
- On-the-fly alignment/re-analysis of SRR accession ids using ebi.ac.uk
- Fast and robust quantification of transcriptome structure and expression using EM
- Dynamic building and entropic measurements of splicing events of any complexity
- Percent-spliced-in (PSI) from event-level EM
- Gene expression (TPM) from transcript-level EM
- Differential splicing comparisons
- Probabilistic calculations of delta PSI leveraging multi-sample biological replicates
Paper: https://doi.org/10.1016/j.molcel.2018.08.018
Get started
1) Install
Whippet v1.6 works on the former long-term support release of Julia (v1.6.7) which is still available here (https://julialang.org). (Note: Whippet.jl does not yet work on Julia v1.9+). If you are new to julia, there is a helpful guide on how to get it up and running here
Download and install dependencies
git clone https://github.com/timbitz/Whippet.jl.git
cd Whippet.jl
julia --project -e 'using Pkg; Pkg.instantiate(); Pkg.test()'
This should tell you Testing Whippet tests passed
Notes:
- Everything in Whippet.jl/bin should work out-of-the-box, however the first time running will be slow as julia will be precompiling code
- Update to the most recent version of Whippet by pulling the master branch
git pull - For all executables in
Whippet.jl/bin, you can use the-hflag to get a list of the available command line options, their usage and defaults. - You should install Julia locally, if you have to install system-wide, there is some help here
- For instructions on using Whippet with Julia v0.6.4, look at the README.md within the Whippet v0.11.1 tag (but please note this verison is no longer supported)
2) Build an index.
a) Annotation (GTF) only index.
You need your genome sequence in fasta, and a gene annotation file in Ensembl-style GTF format. Default GENCODE annotation supplied for hg19 and mm10 in Whippet/anno. You can also obtain Ensembl GTF files from these direct links for Human: Ensembl_hg38_release_92 and Mouse: Ensembl_mm10_release_92. Other Ensembl GTF files can be downloaded here.
Download the genome.
wget https://hgdownload.soe.ucsc.edu/goldenPath/hg19/bigZips/hg19.fa.gz
Build an index.
julia bin/whippet-index.jl --fasta hg19.fa.gz --gtf anno/gencode_hg19.v25.tsl1.gtf.gz
Notes:
- Whippet only uses GTF
exonlines (others are ignored). These must contain bothgene_idandtranscript_idattributes (which should not be the same as one another!). This GTF file should be consistent with the GTF2.2 specification, and should have all entries for a transcript in a continuous block. Warning: The UCSC table browser will not produce valid GTF2.2 files. Similarly, GTF files obtained from iGenomes or the Refseq websites do not satisfy these specifications. - You can specify the output name and location of the index to build using the
-x / --indexparameter. The default (for both whippet-index.jl and whippet-quant.jl) is a generic index namedgraphlocated atWhippet.jl/index/graph.jls, so you must have write-access to this location to use the default.
b) Annotation (GTF) + Alignment (BAM) supplemented index.
Whippet v0.11+ allows you to build an index that includes unannotated splice-sites and exons found in a spliced RNA-seq alignment file. In order to build a BAM supplemented index, you need your BAM file sorted and indexed (using samtools):
# If using multiple BAM files (tissue1, ..., tissue3 etc), merge them first:
samtools merge filename.bam tissue1.bam tissue2.bam tissue3.bam
# If using a single BAM file start here:
samtools sort -o filename.sort filename.bam
samtools rmdup -S filename.sort.bam filename.sort.rmdup.bam
samtools index filename.sort.rmdup.bam
ls filename.sort.rmdup.bam*
filename.sort.rmdup.bam filename.sort.rmdup.bam.bai
Then build an index but with the additional --bam parameter:
julia bin/whippet-index.jl --fasta hg19.fa.gz --bam filename.sort.rmdup.bam --gtf anno/gencode_hg19.v25.tsl1.gtf.gz
Notes:
- The
--bamoption is sensitive to alignment strand, therefore using strand-specific alignments is recommended. - By default only spliced alignments where one of the splice-sites match a known splice-site in the annotation are used, to reduce false positives due to overlapping gene regions (i.e. falsely adding splice sites that belong to a different, but overlapping gene, which is common in many species). Use the
--bam-both-novelflag to override this requirement for greater Recall of unannotated splice-sites. - Control the minimum number of reads required to consider a novel splice site from BAM using the
--bam-min-readsparameter (default is 1). Increase this parameter with large bam files to reduce artifacts and one-off cryptic splice junctions.
3) Quantify FASTQ files.
a) Single-end reads
julia bin/whippet-quant.jl file.fastq.gz
Note: Whippet only accepts standard four-line FASTQ file (described here: https://support.illumina.com/bulletins/2016/04/fastq-files-explained.html)
Also, as of version 1.0.0, --ebi and --url flags have been deprecated to ease maintenance. EBI file paths can be found at the URL http://www.ebi.ac.uk/ena/data/warehouse/filereport?accession=$ebi_id&result=read_run&fields=fastq_ftp&display=txt. Use your own accession id (SRR id) in place of $ebi_id.
For example:
curl "https://www.ebi.ac.uk/ena/data/warehouse/filereport?accession=SRR1199010&result=read_run&fields=fastq_ftp&display=txt"
fastq_ftp
ftp.sra.ebi.ac.uk/vol1/fastq/SRR119/000/SRR1199010/SRR1199010.fastq.gz
b) Paired-end reads
julia bin/whippet-quant.jl fwd_file.fastq.gz rev_file.fastq.gz
To locate paired-end SRR id files, use the same ebi.ac.uk URL:
curl "https://www.ebi.ac.uk/ena/data/warehouse/filereport?accession=ERR1994736&result=read_run&fields=fastq_ftp&display=txt"
fastq_ftp
ftp.sra.ebi.ac.uk/vol1/fastq/ERR199/006/ERR1994736/ERR1994736_1.fastq.gz;ftp.sra.ebi.ac.uk/vol1/fastq/ERR199/006/ERR1994736/ERR1994736_2.fastq.gz
c) Non-default input/output
To specify output location or a specific index:
julia bin/whippet-quant.jl fwd_file.fastq.gz -o outputname -x customindex.jls
You can also output the alignments in SAM format with the --sam flag and convert to bam with samtools:
julia bin/whippet-quant.jl fwd_file.fastq.gz --sam > fwd_file.sam
samtools view -bS fwd_file.sam > fwd_file.bam
For greater stability of quantifications across multiple RNA-seq protocols, try the --biascorrect flag, which will apply GC-content and 5' sequence bias correction methods:
julia bin/whippet-quant.jl fwd_file.fastq.gz --biascorrect
It is also possible to pool fastq files at runtime using shell commands, and the optional (--force-gz) for pooled gz files (files without .gz suffix)
julia bin/whippet-quant.jl <( cat time-series_{1,2,3,4,5}.fastq.gz ) --force-gz -o interval_1-5
4) Compare multiple psi files
Compare .psi.gz files from from two samples -a and -b with any number of replicates (comma delimited list of files or common pattern matching) per sample.
ls *.psi.gz
#sample1-r1.psi.gz sample1-r2.psi.gz sample2-r1.psi.gz sample2-r2.psi.gz
julia bin/whippet-delta.jl -a sample1 -b sample2
#OR
julia bin/whippet-delta.jl -a sample1-r1.psi.gz,sample1-r2.psi.gz -b sample2-r1.psi.gz,sample2-r2.psi.gz
Note: comparisons of single files still need a comma: -a singlefile_a.psi.gz, -b singlefile_b.psi.gz,
Output Formats
The output format for whippet-quant.jl is saved into two core quant filetypes, .psi.gz and .tpm.gz files.
Each .tpm.gz file contains a simple format compatible with many downstream tools (one for the TPM of each annotated transcript, and another at the gene-level):
Gene/Isoform | TpM | Read Counts ---- | --- | ----------- NFIA | 2897.11 | 24657.0
Meanwhile the .psi.gz file is a bit more complex and requires more explanation. Quantified nodes can fall into a number of "node-type" categories:
Type | Interpretation ---- | -------------- CE | Core exon, which may be bounded by one or more alternative AA/AD nodes AA | Alternative Acceptor splice site AD | Alternative Donor splice site RI | Retained intron TS | Tandem tr
Related Skills
node-connect
332.3kDiagnose OpenClaw node connection and pairing failures for Android, iOS, and macOS companion apps
frontend-design
81.7kCreate distinctive, production-grade frontend interfaces with high design quality. Use this skill when the user asks to build web components, pages, or applications. Generates creative, polished code that avoids generic AI aesthetics.
openai-whisper-api
332.3kTranscribe audio via OpenAI Audio Transcriptions API (Whisper).
commit-push-pr
81.7kCommit, push, and open a PR




