Paga
Mapping out the coarse-grained connectivity structures of complex manifolds.
Install / Use
/learn @theislab/PagaREADME
PAGA - partition-based graph abstraction
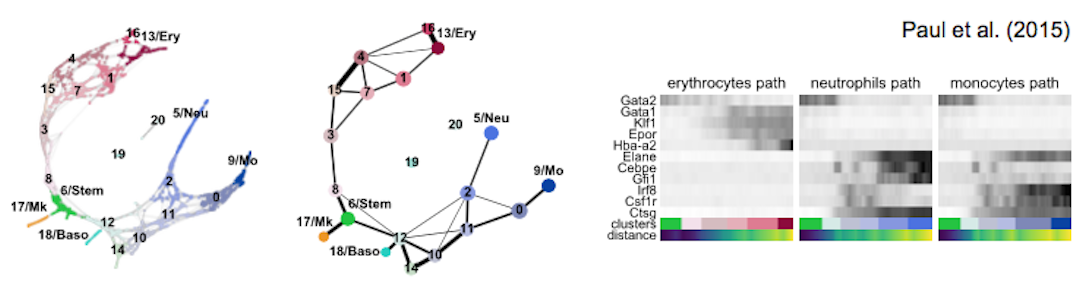
Mapping out the coarse-grained connectivity structures of complex manifolds (Genome Biology, 2019).
PAGA is available within Scanpy through: tl.paga | pl.paga | pl.paga_path | pl.paga_compare.
Below you find links to all central example notebooks, which also allow reproducing all main figures of the paper. If you start working with PAGA, go through blood/paul15.
notebook | system | details | reference | figure ---------------| ---------------| ---------| ----------| ------ blood/simulated | hematopoiesis | simulated | Krumsiek et al., Plos One (2011) | 2a blood/paul15 | murine hematopoiesis | 2,730 cells, MARS-seq | Paul et al., Cell (2015) | 2b blood/nestorowa16 | murine hematopoiesis | 1,654 cells, Smart-seq2 | Nestorowa et al., Blood (2016) | 2c blood/dahlin18 | murine hematopoiesis | 44,802 cells, 10x Genomics | Dahlin et al., Blood (2018) | 2d planaria | planaria | 21,612 cells | Plass et al., Science (2018) | 3 zebrafish | zebrafish embryo | 53,181 cells | Wagner et al., Science (2018) | 4 1M_neurons | neurons | 1.3 million cells, 10x Genomics | 10x Genomics (2017) | S12 deep_learning | cycling Jurkat cells | 30,000 single-cell images | Eulenberg et al., Nat. Commun. (2017) | S14
All supplemental figures of the paper can be reproduced based on the following table.
notebook | description | figure ---------------| ----------| ------ connectivity_measure | connectivity measure | S1, S2, S3 robustness | robustness and multi-resolution capacity | S4, S5 comparisons/simulated_data | comparisons for simulated data | S6, S7 comparisons/paul15_monocle2 | comparison Monocle 2 for Paul et al. (2015) | S8 comparisons/nestorowa16_monocle2 | comparison Monocle 2 for Nestorowa et al. (2016) | S9 embedding_quality | quantifying embedding quality | S10 simulation | simulating hematopoiesis | S11 1M_neurons | neurons, 1.3 million cells, 10x Genomics, 10x Genomics (2017) | S12 blood/paul15 | annotation of louvain clusters using PAGA | S13 deep_learning | cycling Jurkat cells, 30,000 single-cell images, Eulenberg et al., Nat. Commun. (2017) | S14
Related Skills
node-connect
343.1kDiagnose OpenClaw node connection and pairing failures for Android, iOS, and macOS companion apps
frontend-design
90.0kCreate distinctive, production-grade frontend interfaces with high design quality. Use this skill when the user asks to build web components, pages, or applications. Generates creative, polished code that avoids generic AI aesthetics.
openai-whisper-api
343.1kTranscribe audio via OpenAI Audio Transcriptions API (Whisper).
qqbot-media
343.1kQQBot 富媒体收发能力。使用 <qqmedia> 标签,系统根据文件扩展名自动识别类型(图片/语音/视频/文件)。
