Phylosim
An R package for the Monte Carlo simulation of sequence evolution
Install / Use
/learn @sbotond/PhylosimREADME
PhyloSim
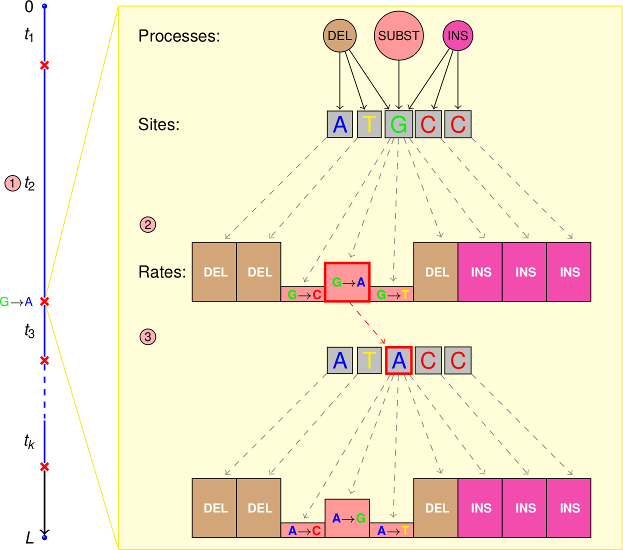
<tt>PhyloSim</tt> is an extensible object-oriented framework for the Monte Carlo simulation of sequence evolution written in 100 percent <tt>R</tt>. It is built on the top of the R.oo and ape packages and uses the Gillespie algorithm to simulate substitutions, insertions and deletions.
<tt>PhyloSim</tt> was brought to you by the Goldman group from EMBL-EBI.
Publication
Botond Sipos, Tim Massingham, Gregory E Jordan and Nick Goldman (2011) <i>PhyloSim - Monte Carlo simulation of sequence evolution in the R statistical computing environment</i> - BMC Bioinformatics 12:104 doi:10.1186/1471-2105-12-104
Download an install
The released packages are available from CRAN.
Key features
-
Simulation of the evolution of a set of discrete characters with arbitrary states evolving by a continuous-time Markov process with an arbitrary rate matrix.
-
Explicit implementations of the most popular substitution models (nucleotide, amino acid and codon substitution models).
-
Simulation under the popular models of among-sites rate variation, like the gamma (<tt>+G</tt>) and invariant sites plus gamma (<tt>+I+G</tt>) models.
-
The possibility to simulate under arbitrarily complex patterns of among-sites rate variation by setting the site specific rates according to any <tt>R</tt> expression.
-
Simulation of one or more separate insertion and/or deletion processes acting on the sequences and which sample the insertion/deletion length from an arbitrary discrete distribution or an <tt>R</tt> expression (so all the probability distributions implemented in <tt>R</tt> are readily available for this purpose).
-
Simulation of the effects of variable functional constraints over the sites by site-process specific insertion and deletion tolerance parameters which determine the rejection probability of a proposed insertion/deletion.
-
The possibility of having a different set of processes and site-process specific parameters for every site, which allows for an arbitrary number of partitions in the simulated data.
-
The possibility to evolve sites by a combination of substitution processes along a single branch.
-
Simulation of heterotachy and other cases of non-homogeneous evolution by allowing the user to set "node hook" functions altering the site properties at internal nodes.
-
The possibility to export the counts of various events ("branch statistics") as <tt>phylo</tt> objects (see the man page of <tt>exportStatTree.PhyloSim</tt>).
-
See the man page of the <tt>PhyloSim</tt> class and the package vignette for more features and examples.
Building from source
The package can be built from the source by issuing <tt>make pack</tt> on a <tt>*nix</tt> system. The building process need the standard unix tools, <tt>Perl</tt> and <tt>R</tt> with the <tt>ape</tt>, <tt>R.oo</tt>, <tt>ggplot2</tt> and <tt>compoisson</tt> packages installed.
Related Skills
node-connect
349.2kDiagnose OpenClaw node connection and pairing failures for Android, iOS, and macOS companion apps
frontend-design
109.5kCreate distinctive, production-grade frontend interfaces with high design quality. Use this skill when the user asks to build web components, pages, or applications. Generates creative, polished code that avoids generic AI aesthetics.
openai-whisper-api
349.2kTranscribe audio via OpenAI Audio Transcriptions API (Whisper).
qqbot-media
349.2kQQBot 富媒体收发能力。使用 <qqmedia> 标签,系统根据文件扩展名自动识别类型(图片/语音/视频/文件)。

