Fastq.bio
An interactive web tool for quality control of DNA sequencing data
Install / Use
/learn @robertaboukhalil/Fastq.bioREADME
fastq.bio
An interactive web tool that generates quality reports from DNA sequencing data without leaving the browser, powered by WebAssembly.
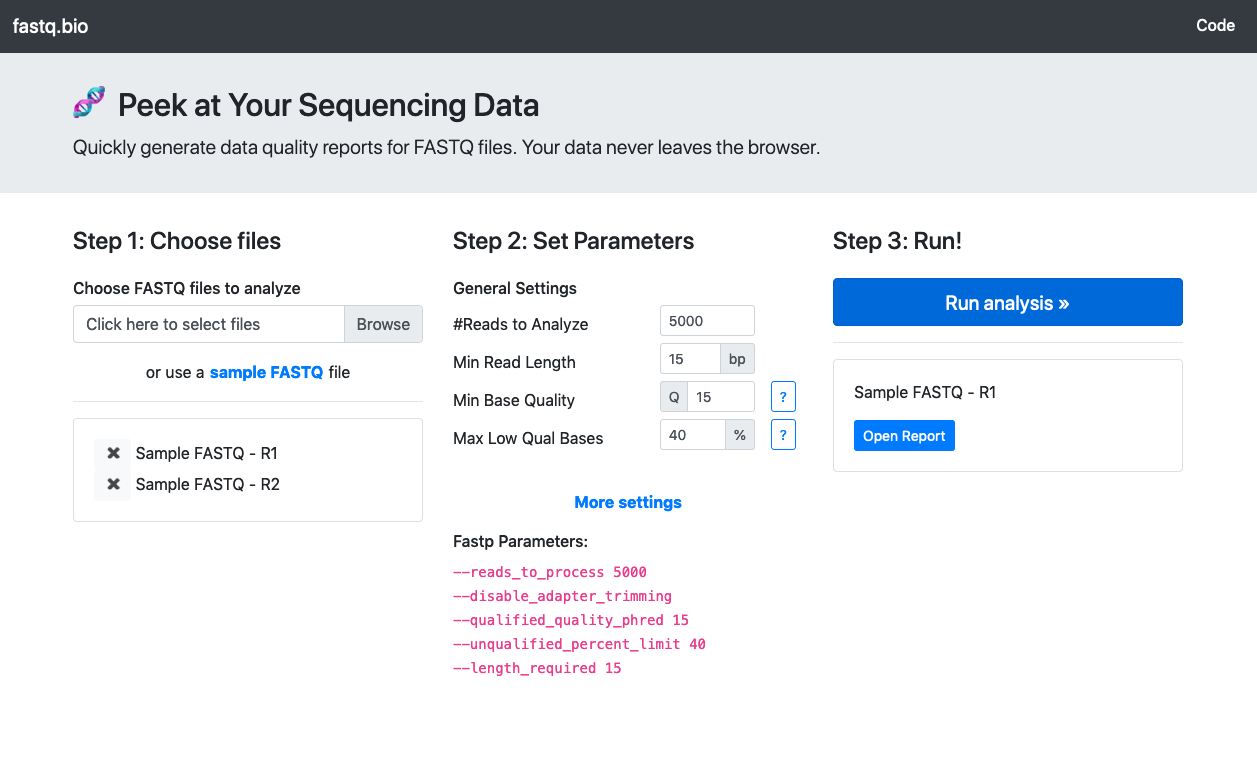
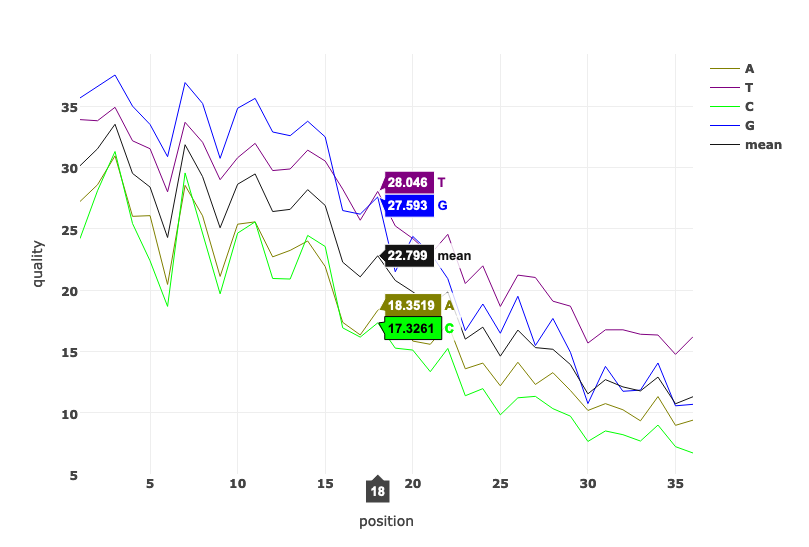
How to use fastq.bio
You can use the hosted version at fastq.bio.
Or, to run fastq.bio locally:
npm install
npm run dev
and open the URL specified on the command line (e.g. http://localhost:5000).
How it works
- To generate the QC report, fastq.bio runs the C tool fastp directly in the browser using WebAssembly. For details about the compilation from C to WebAssembly, see the biowasm project.
- fastq.bio uses the aioli library to run the WebAssembly module in a WebWorker, and handles mounting user files to a virtual file system.
- For details about how WebAssembly can in some cases be a powerful tool for speeding up web apps, see my Smashing Magazine article.
Related Skills
node-connect
343.3kDiagnose OpenClaw node connection and pairing failures for Android, iOS, and macOS companion apps
frontend-design
92.1kCreate distinctive, production-grade frontend interfaces with high design quality. Use this skill when the user asks to build web components, pages, or applications. Generates creative, polished code that avoids generic AI aesthetics.
openai-whisper-api
343.3kTranscribe audio via OpenAI Audio Transcriptions API (Whisper).
qqbot-media
343.3kQQBot 富媒体收发能力。使用 <qqmedia> 标签,系统根据文件扩展名自动识别类型(图片/语音/视频/文件)。
