Nxontology
NetworkX-based Python library for representing ontologies
Install / Use
/learn @related-sciences/NxontologyREADME
NetworkX-based Python library for representing ontologies
Summary
nxontology is a Python library for representing ontologies using a NetworkX graph. Currently, the main area of functionality is computing similarity measures between pairs of nodes.
Usage
Here, we'll use the example metals ontology:
Note that NXOntology represents the ontology as a networkx.DiGraph, where edge direction goes from superterm to subterm.
Given an NXOntology instance, here how to compute intrinsic similarity metrics.
from nxontology.examples import create_metal_nxo
metals = create_metal_nxo()
# Freezing the ontology prevents adding or removing nodes or edges.
# Frozen ontologies cache expensive computations.
metals.freeze()
# Get object for computing similarity, using the Sanchez et al metric for information content.
similarity = metals.similarity("gold", "silver", ic_metric="intrinsic_ic_sanchez")
# Access a single similarity metric
similarity.lin
# Access all similarity metrics
similarity.results()
The final line outputs a dictionary like:
{
'node_0': 'gold',
'node_1': 'silver',
'node_0_subsumes_1': False,
'node_1_subsumes_0': False,
'n_common_ancestors': 3,
'n_union_ancestors': 5,
'batet': 0.6,
'batet_log': 0.5693234419266069,
'ic_metric': 'intrinsic_ic_sanchez',
'mica': 'coinage',
'resnik': 0.8754687373538999,
'resnik_scaled': 0.48860840553061435,
'lin': 0.5581154235118403,
'jiang': 0.41905978419640516,
'jiang_seco': 0.6131471927654584,
}
It's also possible to visualize the similarity between two nodes like:
from nxontology.viz import create_similarity_graphviz
gviz = create_similarity_graphviz(
# similarity instance from above
similarity,
# show all nodes (defaults to union of ancestors)
nodes=list(metals.graph),
)
# draw to PNG file
gviz.draw("metals-sim-gold-silver-all.png"))
Resulting in the following figure:
<!-- from test output: cp nxontology/tests/viz_outputs/metals-sim-gold-silver-all.png media/ -->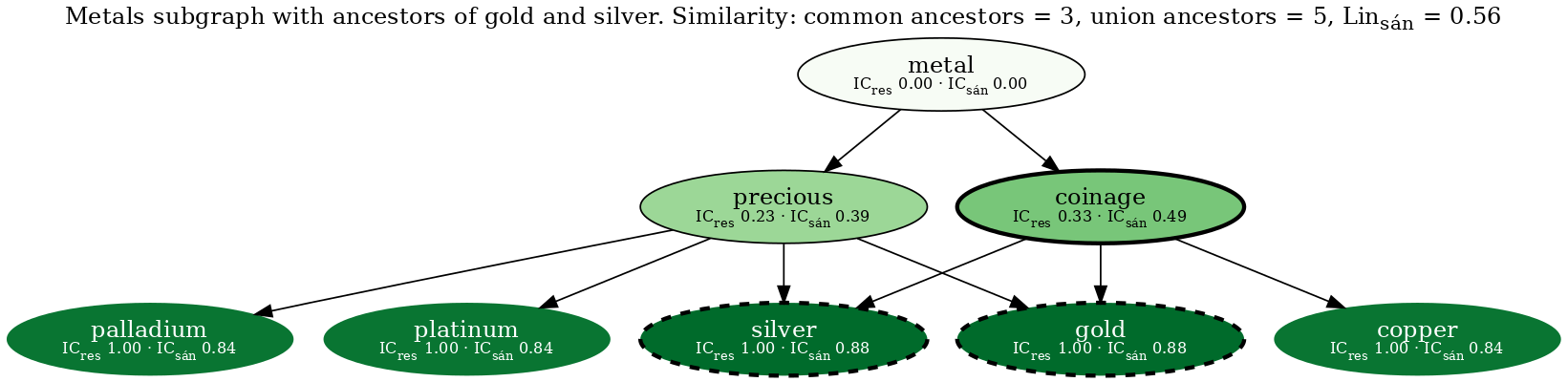
The two query nodes (gold & silver) are outlined with a bold dashed line. Node fill color corresponds to the Sánchez information content, such that darker nodes have higher IC. The most informative common ancestor (coinage) is outlined with a bold solid line. Nodes that are not an ancestor of gold or silver have an invisible outline.
Loading ontologies
Pronto supports reading ontologies from the following file formats:
- Open Biomedical Ontologies 1.4:
.oboextension, uses the fastobo parser. - OBO Graphs JSON:
.jsonextension, uses the fastobo parser. - Ontology Web Language 2 RDF/XML:
.owlextension, uses the prontoRdfXMLParser.
The files can be local or at a network location (URL starting with https, http, or ftp). Pronto detects and handles gzip, bzip2, and xz compression.
Here are examples operations on the Gene Ontology, using pronto to load the ontology:
>>> from nxontology.imports import from_file
>>> # versioned URL for the Gene Ontology
>>> url = "http://release.geneontology.org/2021-02-01/ontology/go-basic.json.gz"
>>> nxo = from_file(url)
>>> nxo.n_nodes
44085
>>> # similarity between "myelination" and "neurogenesis"
>>> sim = nxo.similarity("GO:0042552", "GO:0022008")
>>> round(sim.lin, 2)
0.21
>>> import networkx as nx
>>> # Gene Ontology domains are disconnected, expect 3 components
>>> nx.number_weakly_connected_components(nxo.graph)
3
>>> # Note however that the default from_file reader only uses "is a" relationships.
>>> # We can preserve all GO relationship types as follows
>>> from collections import Counter
>>> import pronto
>>> from nxontology import NXOntology
>>> from nxontology.imports import pronto_to_multidigraph, multidigraph_to_digraph
>>> go_pronto = pronto.Ontology(handle=url)
>>> go_multidigraph = pronto_to_multidigraph(go_pronto)
>>> Counter(key for _, _, key in go_multidigraph.edges(keys=True))
Counter({'is a': 71509,
'part of': 7187,
'regulates': 3216,
'negatively regulates': 2768,
'positively regulates': 2756})
>>> go_digraph = multidigraph_to_digraph(go_multidigraph, reduce=True)
>>> go_nxo = NXOntology(go_digraph)
>>> # Notice the similarity increases due to the full set of edges
>>> round(go_nxo.similarity("GO:0042552", "GO:0022008").lin, 3)
0.699
>>> # Note that there is also a dedicated reader for the Gene Ontology
>>> from nxontology.imports import read_gene_ontology
>>> read_gene_ontology(release="2021-02-01")
Users can also create their own networkx.DiGraph to use this package.
Prebuilt Ontologies
The nxontology-data repository creates NXOntology objects for many popular ontologies / taxonomies.
Installation
nxontology can be installed with pip from PyPI like:
# standard installation
pip install nxontology
# installation with viz extras
pip install nxontology[viz]
The extra viz dependencies are required for the nxontology.viz module.
This includes pygraphviz, which requires a pre-existing graphviz installation.
Development
Some helpful development commands:
# create a virtual environment for development
python3 -m venv .venv
# activate virtual environment
source .venv/bin/activate
# install package for development
pip install --editable ".[dev,viz]"
# Set up the git pre-commit hooks.
# `git commit` will now trigger automatic checks including linting.
pre-commit install
# Run all pre-commit checks (CI will also run this).
pre-commit run --all
# run tests
pytest
Releases are created on GitHub.
The release action defined by release.yaml will build the distribution and upload to PyPI.
The package version is automatically generated from the git tag by setuptools_scm.
Bibliography
Here's a list of alternative projects with code for computing semantic similarity measures on ontologies:
- Ontology Access Kit (OAK) in Python.
- Semantic Measures Library & ToolKit at sharispe/slib in Java.
- DiShIn at lasigeBioTM/DiShIn in Python.
- Sematch at gsi-upm/sematch in Python.
- ontologySimilarity mirrored at cran/ontologySimilarity. Part of the ontologyX suite of R packages.
- Materials for Machine Learning with Ontologies at bio-ontology-research-group/machine-learning-with-ontologies (compilation)
Below are a list of references related to ontology-derived measures of similarity.
Feel free to add any reference that provides useful context and details for algorithms supported by this package.
Metadata for a reference can be generated like manubot cite --yml doi:10.1016/j.jbi.2011.03.013.
Adding CSL YAML output to media/bibliography.yaml will cache the metadata and allow manual edits in case of errors.





