Reseek
Protein structure alignment and search algorithm
Install / Use
/learn @rcedgar/ReseekREADME

Reseek is a protein structure search and alignment algorithm which improves sensitivity in protein homolog detection compared to state-of-the-art methods including DALI, TM-align and Foldseek with similar speed to Foldseek.
Online structure search
Search a protein structure against AFDB, PDB or BFVD with typical results in 2 to 5 minutes.
<hr> <hr>Reseek achieves highest accuracy in homolog detection and E-values
On the SCOP40 benchmark test (see results later below), Reseek has substantially higher ability to discriminate homologs compared to previous algorithms including DALI, TM-align and Foldseek. This means that Reseek is better at sorting true homologs ahead of false positives.
Reseek also provides a much more accurate estimate of statistical significance (E-value), enabling users to set a cutoff based on an acceptable number of false positives for a given search, while DALI and Foldseek often over-estimate significance by 5 to 6 orders of magnitude (references below).
YouTube talk describing the algorithm
Reseek is based on sequence alignment where each residue in the protein backbone is represented by a letter in a novel “mega-alphabet” of 85,899,345,920 (∼10<sup>11</sup>) distinct structure states. This talk explains how it works.
<img src="https://drive5.com/reseek/youtube_snip.gif" width="150">
Command line
<pre> Common commands -search # Alignment (e.g. DB search, pairwise, all-vs-all) -convert # Convert file formats (e.g. create DB) -alignpair # Pair-wise alignment and superposition Search against database reseek -search STRUCTS -db STRUCTS -output hits.txt # STRUCTS specifies structure(s), see below Recommended format for large database is .bca, e.g. reseek -convert /data/PDB_mirror/ -bca PDB.bca Align and superpose two structures reseek -alignpair 1XYZ.pdb -input2 2ABC.pdb -aln FILE # Sequence alignment (text) -output FILE # Rotated 1XYZ (PDB format) All-vs-all alignment reseek -search STRUCTS -output hits.txt Output options for -search -aln FILE # Alignments in human-readable format -output FILE # Hits in tabbed text format -columns name1+name2+name3... # Output columns, names are # query Query label # target Target label # qlo Start of aligment in query # qhi End of aligment in query # tlo Start of aligment in target # thi End of aligment in target # ql Query length # tl Target length # pctid Percent identity of alignment # cigar CIGAR string # pvalue P-value according to log-linear null model (RECOMMENDED) # evalue E-value according to log-linear null model (DEPRECATED) # aq AQ (aln. qual., 0 to 1) (DEPRECATED) # qrow Aligned query sequence with gaps (local) # trow Aligned target sequence with gaps (local) # qrowg Aligned query sequence with gaps (global) # trowg Aligned target sequence with gaps (global) # std query+target+qlo+qhi+ql+tlo+thi+tl+pctid+pvalue (default) Search and alignment options -fast, -sensitive or -verysensitive # Required -evalue E # Max E-value (default 10 unless -verysensitive) -omega X # Omega accelerator (floating-point) -minu U # K-mer accelerator (integer) -gapopen X # Gap-open penalty (floating-point >= 0) -gapext X # Gap-extend penalty (floating-point >= 0) -dbsize D # DB size (nr. chains) for E-value (default actual size) Convert between file formats reseek -convert STRUCTS [one or more output options] -cal FILENAME # .cal format, text with a.a. and C-alpha x,y,z -bca FILENAME # .bca format, binary .cal, recommended for DBs -fasta FILENAME # FASTA format Create input for Muscle-3D multiple structure alignment: reseek -pdb2mega STRUCTS -output structs.mega STRUCTS argument is one of: NAME.cif or NAME.mmcif # PDBx/mmCIF file NAME.pdb # Legacy format PDB file NAME.cal # C-alpha tabbed text format with chain(s) NAME.bca # Binary C-alpha, recommended for larger DBs NAME.files # Text file with one STRUCT per line, # may be filename, directory or .files DIRECTORYNAME # Directory (and its sub-directories) is searched # for known file types including .pdb, .files etc. Other options: -log FILENAME # Log file with errors, warnings, time and memory. -threads N # Number of threads, default number of CPU cores. </pre>Build from source on Linux x86
<pre> cd src/; chmod +x build_linux_x86.bash ; ./build_linux_x86.bash </pre>Build from source on Windows
Load reseek.vcxproj into Microsoft Visual Studio and use the Build command.
OSX currently not supported
The problem is compatibility with the amazing parasail library https://github.com/jeffdaily/parasail (thanks Jeff!) which reseek uses for fast Smith-Waterman alignment. See issue 25, there is probably an easy fix, anyone...?
Ignore static link warning
Don't worry about a warning something like this, it's expected:
<pre> warning: Using 'dlopen' in statically linked applications requires at runtime the shared libraries from the glibc version used for linking </pre>More documentation
SCOP40 benchmark code and results
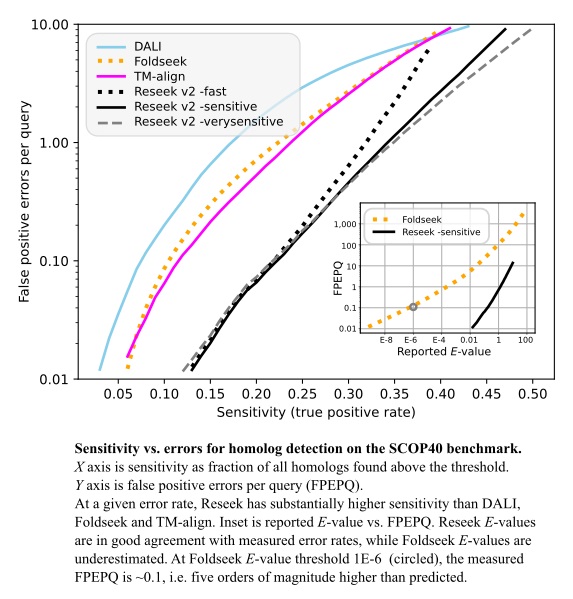
Method sensitivity was measured on the SCOP40 benchmark using superfamily as the truth standard, focusing on the regime with false-positive error rates <10 per query, corresponding to E<10 for an ideal E-value.
https://github.com/rcedgar/reseek_bench
References
Edgar RC. "Protein structure alignment by Reseek improves sensitivity to remote homologs" (Bioinformatics 2024) Nov;40(11):btae687. https://academic.oup.com/bioinformatics/article/40/11/btae687/7901215
Edgar RC. and Sahakyan S. "Protein structure alignment significance is often exaggerated" (bioRxiv 2025) https://www.biorxiv.org/content/10.1101/2025.07.17.665375v1
