Ipcoal
Simulation package for getting gene tree distributions and loci / SNPs from input tree topologies and migration / Ne scenarios. Wraps around msprime.
Install / Use
/learn @pmckenz1/IpcoalREADME
ipcoal
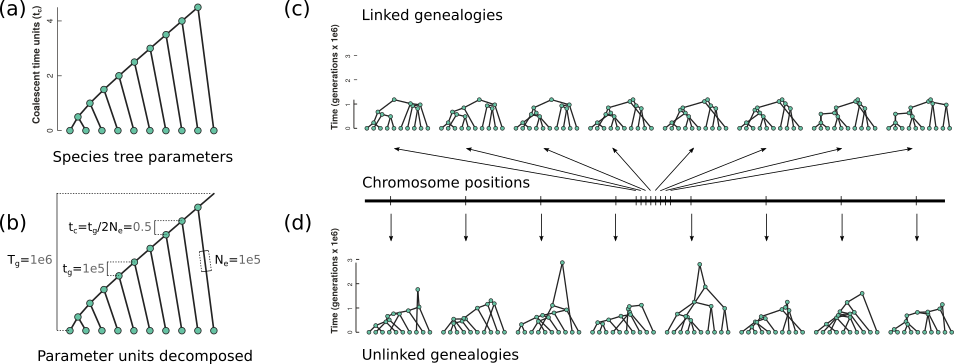
Python package to interactively simulate genealogies and sequence data under the multispecies coalescent
Full documentation here: https://ipcoal.readthedocs.io
import ipcoal
import toytree
# get a species tree
tree = toytree.rtree.baltree(ntips=6, treeheight=1e6)
# init a simulator from the tree w/ additional parameters
model = ipcoal.Model(tree, Ne=1e6)
# simulate genealogies and sequences
model.sim_loci(nloci=5, nsites=1e5)
# access results in a dataframe
model.df
# infer gene trees for each locus, or in sliding windows
model.infer_gene_trees()
# write sequences to a file
model.write_loci_to_phylip()