C302
The c302 framework for generating multiscale network models of C. elegans
Install / Use
/learn @openworm/C302README
The c302 modelling framework for C. elegans
c302 is a framework for generating network models in NeuroML 2 based on C. elegans connectivity data. It is primarily intended as a way to generate neuronal networks at multiple levels of detail for the OpenWorm project. The c302 framework is described in the following publication:
c302: a multiscale framework for modelling the nervous system of Caenorhabditis elegans Padraig Gleeson, David Lung, Radu Grosu, Ramin Hasani, Stephen D. Larson, Phil. Trans. R. Soc. B 2018 373 20170379; DOI: 10.1098/rstb.2017.0379.
To see how c302 relates to other OpenWorm subprojects click on the image below:
<p align="center"><a href="OpenWormOverview.md"><img width="200" alt="c302_in_overview" src="https://user-images.githubusercontent.com/1573896/44876490-42fca100-ac6f-11e8-95ab-76dc8a2d80e1.png"></a></p>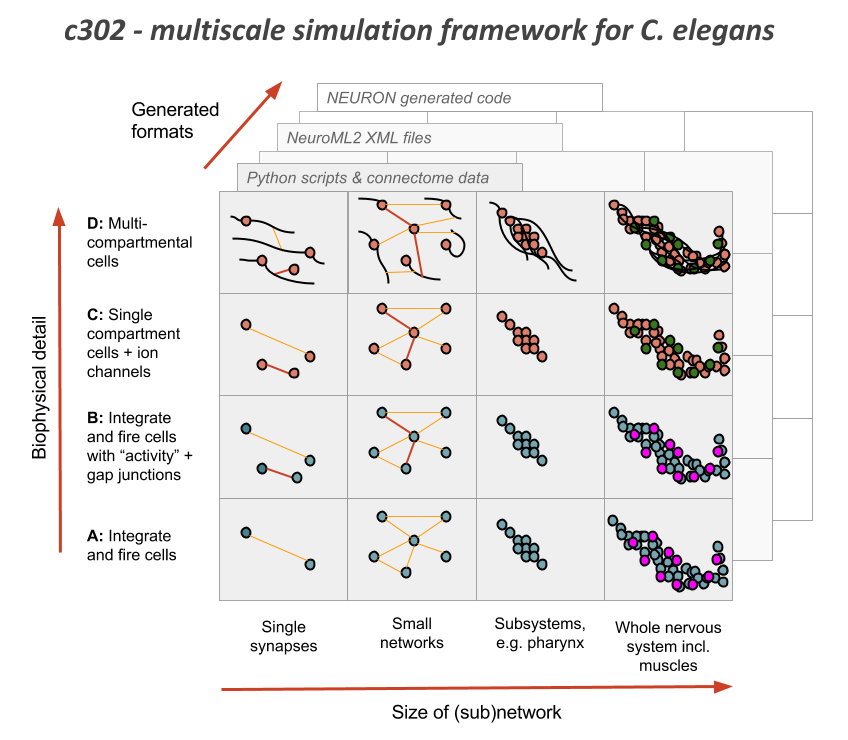
It uses information on the synaptic connectivity of the network (from here) and uses libNeuroML to generate a network in valid NeuroML, which can be run in jNeuroML or pyNeuroML.
To install & test
The full set of dependencies for c302 can be installed with the following (see also the ci.yml and non_omv.yml workflow files):
git clone https://github.com/openworm/c302.git
cd c302
pip install .
# The following can only be run on Mac and Linux. Note: c302 can be run without ownmeta fully installed
owm bundle remote --user add ow 'https://raw.githubusercontent.com/openworm/owmeta-bundles/master/index.json'
This will install c302 as well as pyNeuroML and owmeta.
For Windows users a virtual environment must be generated with Python 3.10 as any newer versions do not currently support owmeta.
To run the test.sh script, the Neuron simulator is also required, and should be installed as described here (On Mac and Linux it can be installed with pip install neuron).
For Linux users, additional dependencies (OpenJDK 19 and Graphviz) are required to run the usage examples:
sudo apt-get install openjdk-19-jdk graphviz
Please note that Python3 is also required to run both the test.sh script and the Usage Examples. For convenience on Linux, it is recommended to install python-is-python3 prior to running the test.sh script.
sudo apt-get install python-is-python3
Quick test
To ensure everything is set up correctly try:
-
Regenerate the NeuroML & LEMS files for one instance of the model:
python c302/c302_Pharyngeal.py B # generate pharyngeal network (see fig. above) using parameter set B -
Run a simulation with pyNeuroML:
pynml examples/LEMS_c302_B_Pharyngeal.xml
To test all of the working features of the framework run test.sh:
./test.sh
Docker image
c302 is included in the Docker image which contains the complete OpenWorm simulation stack. Click on the image below for more details.
<p align="center"><a href="https://github.com/openworm/OpenWorm/blob/master/README.md#quickstart"><img width="400" alt="Worm Crawling" src="https://raw.githubusercontent.com/openworm/OpenWorm/master/img/worm-crawling.gif"></a></p>Usage Examples
1a) Run standard examples with pyNeuroML
There are a number of example models included with the standard distribution. These consist of: A) generated NeuroML 2 network description file (example), containing the definitions of the cells to use (e.g. iafCell for an integrate and fire cell), any inputs (e.g. pulseGenerator) as well as the populations, projections and inputLists contained within the network (for a full description of the NeuroML elements see here); and B) a LEMS simulation file (example) describing how long to simulate, the timestep and what to plot/record.
# generate 2 neurons & 1 muscle with current inputs using parameter set A
pynml examples/LEMS_c302_A_IClamp.xml
# generate full scale network using parameter set C
pynml examples/LEMS_c302_C_Full.xml
# generate pharyngeal network using parameter set B
pynml examples/LEMS_c302_B_Pharyngeal.xml
Screenshots of a simulation with pyNeuroML of c302_B_Pharyngeal are shown below (left: membrane potential of 20 cells, right: "activity" of 20 cells - a value from 0-1 showing time smoothed activity of each cell):
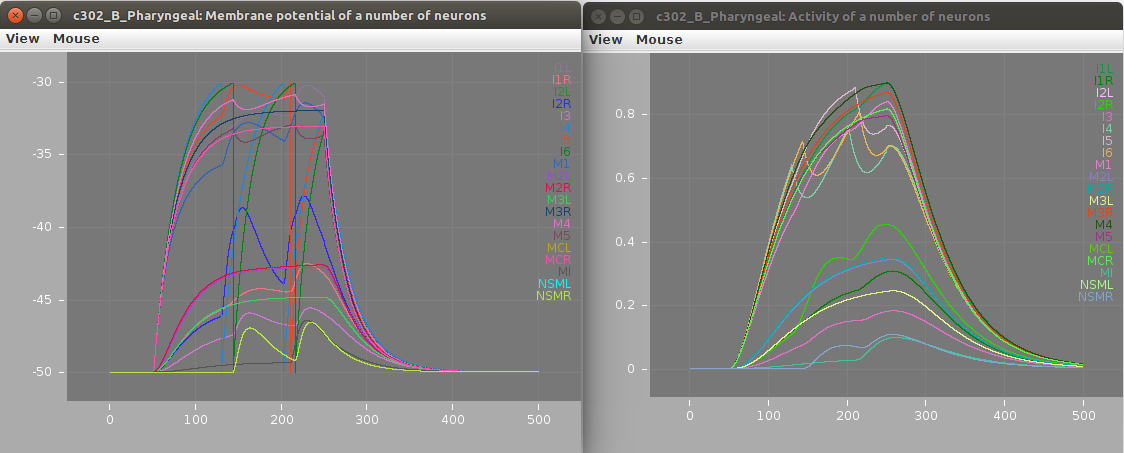
1b) Run standard examples with Neuron
The models can also be run using the Neuron simulator. This should be installed as outlined here. Note: running c302 with Neuron is not currently supported on Windows.
cd examples
pynml LEMS_c302_A_IClamp.xml -neuron # Generate the Neuron files (Python/hoc/mod)
nrnivmodl # Compile the mod files (used for cell/ion channel definitions)
nrngui LEMS_c302_A_IClamp_nrn.py # Run the main Python file for the simulation using Neuron
Note: models with the D parameter set can only be run using Neuron (not pyNeuroML), simnce they consist of multicompartmental Neurons, e.g.
pynml LEMS_c302_D_Pharyngeal.xml -neuron
nrnivmodl
nrngui LEMS_c302_D_Pharyngeal_nrn.py
This produces the following (graph on top is [Ca2+], bottom is membrane potential; 3D view on right can be produced by selecting in the Neuron main menu: Graph -> Shape plot)
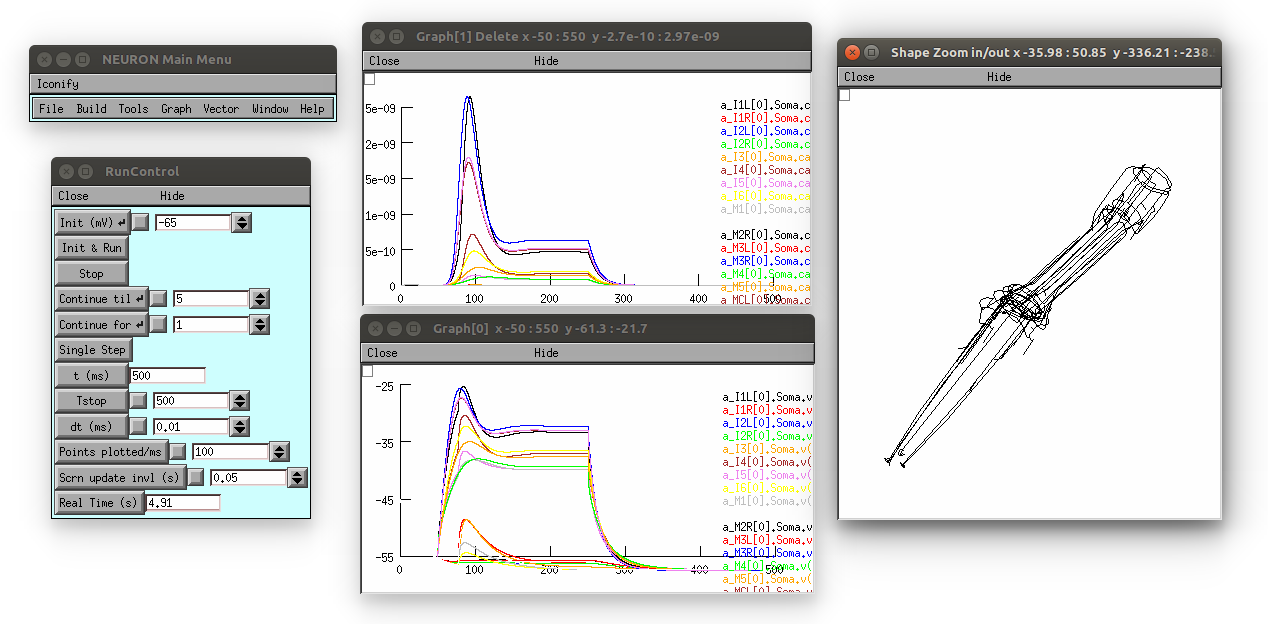
2) Use command line interface to create new network
The c302 command line utility can be used to generate customised networks of varying size, with different cells stimulated, of varying duration from the command line:
c302 MyNetwork parameters_C -cells ["AVBR","VD3"] -cellstostimulate ["AVBR"] -paramoverride {"unphysiological_offset_current":"2.9pA"} -duration 300
This will create a NeuroML 2 file and a LEMS file to execute it, containing 2 cells, stimulating 1 of them, and with a duration of 300 ms. It can be run with:
pynml LEMS_MyNetwork.xml
This produces two plots: One is neuron activity, measured as calcium ion concentration. You will likely need to click "Frame" in the "View" menu to scale the plot.

The other shows membrane potential for each neuron in the network.

To see the structure of the network, use pyNeuroML (please note that Graphviz is required):
pynml MyNetwork.net.nml -graph 4c # Try other options like 1, 2f, 5c for varying levels of detail

More options for using the c302 command can be found with
c302 -h
3) Change parameters in a file
To investigate how the behaviour of a model changes when parameters are varied, it is possible to change the parameters in the parameters_X.py files and regenerate.
For example in parameters_C.py there are lists of parameters like:
self.add_bioparameter("muscle_leak_cond_density", "5e-7 S_per_cm2", "BlindGuess", "0.1")
self.add_bioparameter("neuron_leak_cond_density", "0.005 mS_per_cm2", "BlindGuess", "0.1")
self.add_bioparameter("leak_erev", "-50 mV", "BlindGuess", "0.1")
To change the model behaviour alter one of these values, e.g.
self.add_bioparameter("neuron_leak_cond_density", "0.02 mS_per_cm2", "BlindGuess", "0.1")
and look at the behaviour afterwards (note the package needs to be reinstalled)
sudo python setup.py install # reinstall package after change
python c302/c302_IClamp.py C # regenerate c302_C_IClamp
pynml examples/LEMS_c302_C_IClamp.xml # run simulation
The plots below show the neuron's membrane potential on application of 6 increasing pulses of current before (left) and after (right) the change, indicating how increasing the leak conductance removes the spiking:
<p><img src="https://raw.githubusercontent.com/openworm/c302/master/images/changePre.png" width=400/> <img src="https://raw.githubusercontent.com/openworm/c302/master/images/changePost.png" width=400/></p>4) Adding a new input type to NeuroML model
The structure of the model generated can be altered by modifying the NeuroML model returned in the c302_XXX.py script. As an example, say we want to add a sine wave current to the Muscles network (specified by [c302_Muscles.py](https://github.com/openworm/c302/
