RVAgene
Recurrent Variational Auto gene encoder
Install / Use
/learn @maclean-lab/RVAgeneREADME
RVAgene: modeling gene expression dynamics
Overview
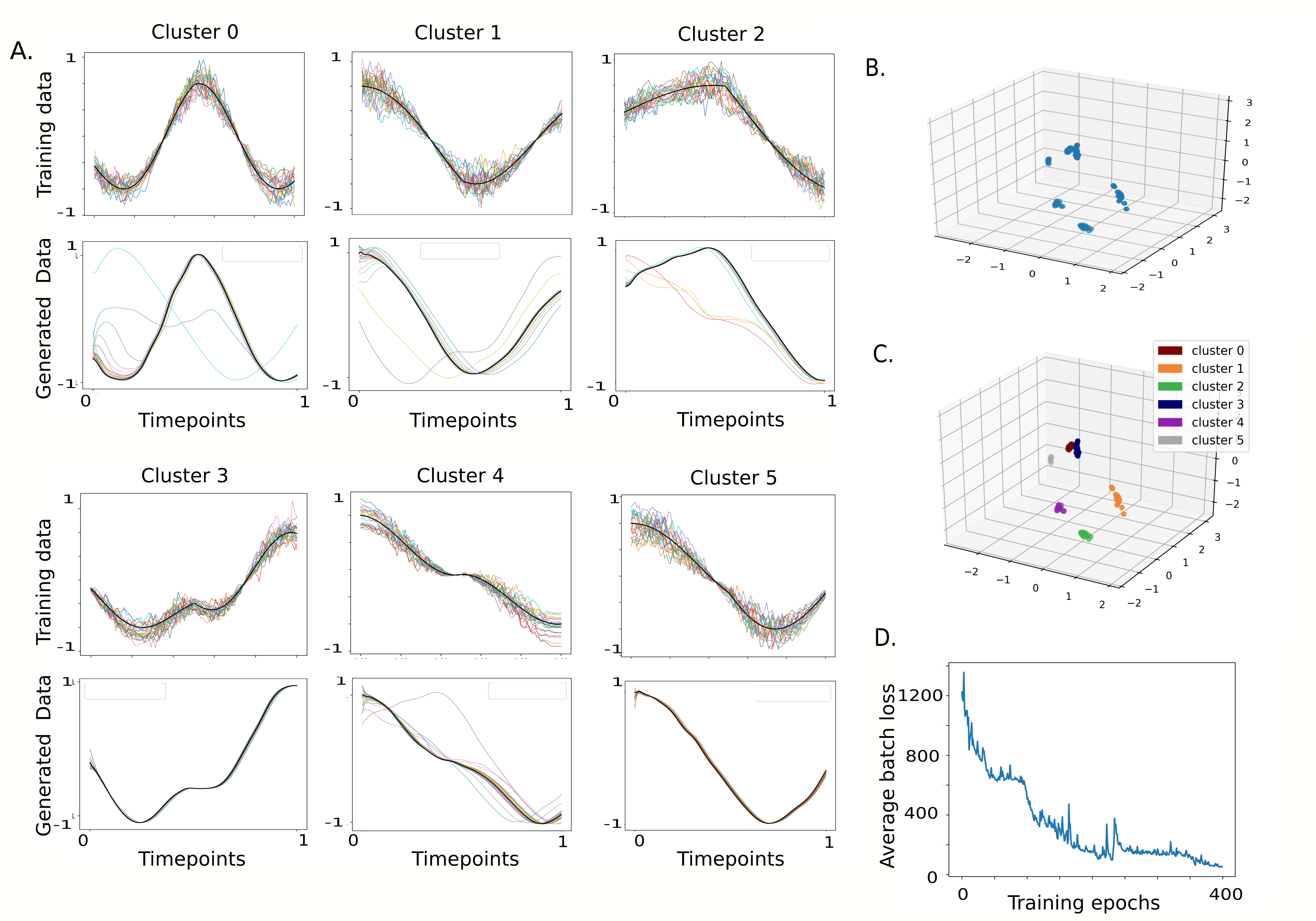
RVAgene models gene expression dynamics in single-cell or bulk data. Read the paper here.
Requirements
- Python 3
- numpy, matplotlib, pytorch, scikit-learn, tqdm
- GPU (optional)
Quickstart
- Jupyter notebook demo of RVAgene here
- Jupyter notebook demo of processing bulk Single cell data and running RVAgene here
- Or on the command line
- python gen_synthetic_data.py <dataset_name> e.g.
python gen_synthetic_data.py demosim - python train_and_gen.py <dataset_name> e.g.
python train_and_gen.py demosim
- python gen_synthetic_data.py <dataset_name> e.g.
Contents
data : contains example synthetic gene expression time series data with 6 inherent clusters <br />
rvagene : contains code for recurrent variational autoencoder <br />
train_and_gen.py : code demonstrating training RVAgene, unsupervised clustering on latent space using K-means and Generating new gene expression data by sampling and decoding points from latent space. <br />
gen_synthetic_data.py : code to generate synthetic data with cluster structure as described in the paper. <br />
figs : contains figures generated by the demo code. <br />
demo.ipynb : Demonstration on the whole synthetic data generation, RVAgene training, clustering on latent space and new cluster specific data generation process <br />
single_cell_demo.ipynb :demo of processing bulk Single cell data and running RVAgene
Model and training parameters:
sequence_length: length of the input sequence <br />
number_of_features : number of features per timepoint per gene i.e. 1 <br />
hidden_size: hidden size of the RNN <br />
hidden_layer_depth: number of layers in RNN (1 is enough) <br />
latent_length: latent vector length <br />
batch_size: number of genes in a single batch. IMPORTANT: last batch will be dropped is it is not a divisor of number of training genes. <br />
learning_rate: the learning rate of the module <br />
n_epochs: Number of iterations/epochs <br />
dropout_rate: The probability of a node being dropped-out <br />
optimizer: ADAM/ SGD optimizer to reduce the loss function <br />
loss: SmoothL1Loss / MSELoss / ReconLoss / any custom loss which inherits from _Loss class <br />
boolean cuda: to be run on GPU or not <br />
print_every: The number of iterations after which loss should be printed for each epoch <br />
boolean clip: Gradient clipping to overcome explosion <br />
max_grad_norm: The grad-norm to be clipped if using clipping <br />
dload: Download directory where models are to be dumped <br />
log_file: File to log training loss , default None i.e. STDOUT <br />
Datasets
scRNA seq Data used in the paper from : https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE65525 (Klein et. al. 2015) Bulk RNA seq Data used in the paper from : https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE98622 (Liu. et. al. , 2017)
Acknowledgments
- Thanks to open source implementation of recurrent VAE at https://github.com/tejaslodaya/timeseries-clustering-vae
- Relevant research works as cited in the work.
Contributors
<a href="https://github.com/maclean-lab/RVAgene/graphs/contributors"> <img src="https://contributors-img.web.app/image?repo=maclean-lab/RVAgene" /> </a>Related Skills
node-connect
353.3kDiagnose OpenClaw node connection and pairing failures for Android, iOS, and macOS companion apps
frontend-design
111.7kCreate distinctive, production-grade frontend interfaces with high design quality. Use this skill when the user asks to build web components, pages, or applications. Generates creative, polished code that avoids generic AI aesthetics.
openai-whisper-api
353.3kTranscribe audio via OpenAI Audio Transcriptions API (Whisper).
qqbot-media
353.3kQQBot 富媒体收发能力。使用 <qqmedia> 标签,系统根据文件扩展名自动识别类型(图片/语音/视频/文件)。
