ADCME.jl
Automatic Differentiation Library for Computational and Mathematical Engineering
Install / Use
/learn @kailaix/ADCME.jlREADME
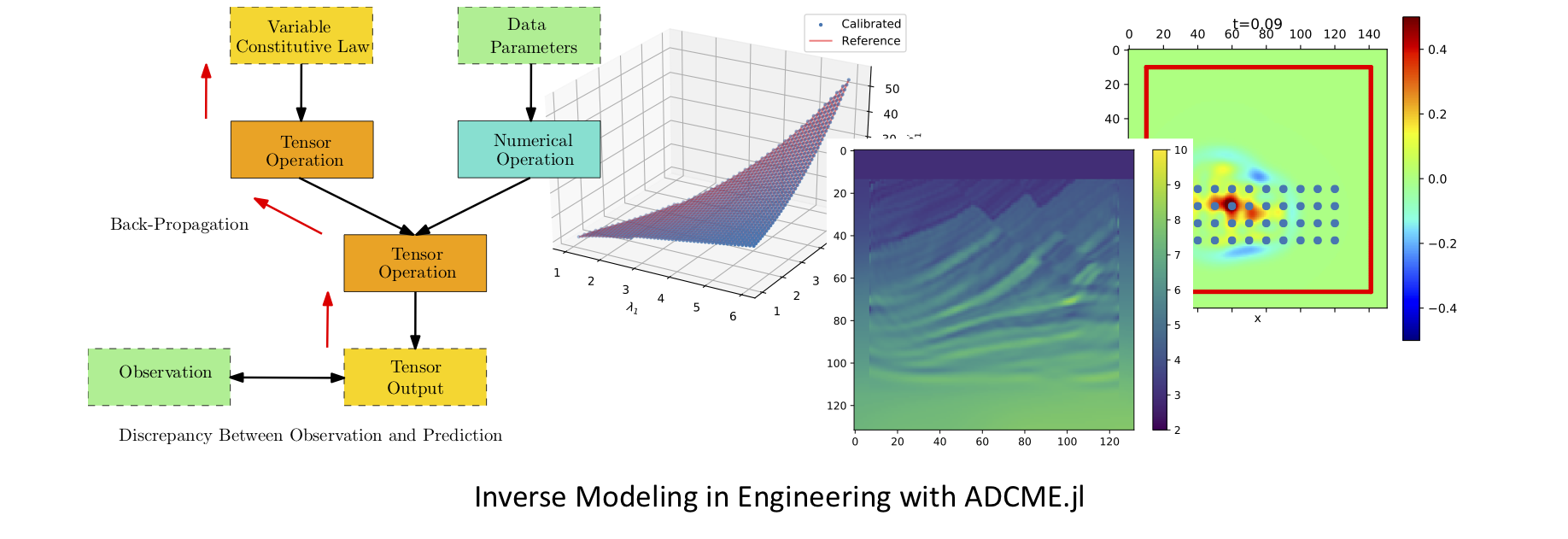
The ADCME library (Automatic Differentiation Library for Computational and Mathematical Engineering) aims at general and scalable inverse modeling in scientific computing with gradient-based optimization techniques. It is built on the deep learning framework, graph-mode TensorFlow, which provides the automatic differentiation and parallel computing backend. The dataflow model adopted by the framework makes it suitable for high performance computing and inverse modeling in scientific computing. The design principles and methodologies are summarized in the slides.
Check out more about slides and videos on ADCME!
| Install ADCME and Get Started (Windows, Mac, and Linux) | Scientific Machine Learning for Inverse Modeling | Solving Inverse Modeling Problems with ADCME | ...more on ADCME Youtube Channel! |
| ------------------------------------------------------------ | ------------------------------------------------------------ | ------------------------------------------------------------ | ------------------------------------------------------------ |
|  |
|  |
|  |
| |
Several features of the library are
- MATLAB-style Syntax. Write
A*Bfor matrix production instead oftf.matmul(A,B). - Custom Operators. Implement operators in C/C++ for performance critical parts; incorporate legacy code or specially designed C/C++ code in
ADCME; automatic differentiation through implicit schemes and iterative solvers. - Numerical Scheme. Easy to implement numerical schemes for solving PDEs.
- Physics Constrained Learning. Embed neural network into PDEs and solve with any numerical schemes, including implicit and iterative schemes.
- Static Graphs. Compilation time computational graph optimization; automatic parallelism for your simulation codes.
- Parallel Computing. Concurrent execution and model/data parallel distributed optimization.
- Custom Optimizers. Large scale constrained optimization? Use
CustomOptimizerto integrate your favorite optimizer. Try out prebuilt Ipopt and NLopt optimizers. - Sparse Linear Algebra. Sparse linear algebra library tailored for scientific computing.
- Inverse Modeling. Many inverse modeling algorithms have been developed and implemented in ADCME, with wide applications in solid mechanics, fluid dynamics, geophysics, and stochastic processes.
Finite Element Method. Get AdFem.jl today for finite element simulation and inverse modeling!
Start building your forward and inverse modeling using ADCME today!
| Documentation | Tutorial | Applications |
| ------------------------------------------------------------ | ------------------------------------------------------------ | ------------------------------------------------------------ |
| 


Graph-mode TensorFlow for High Performance Scientific Computing
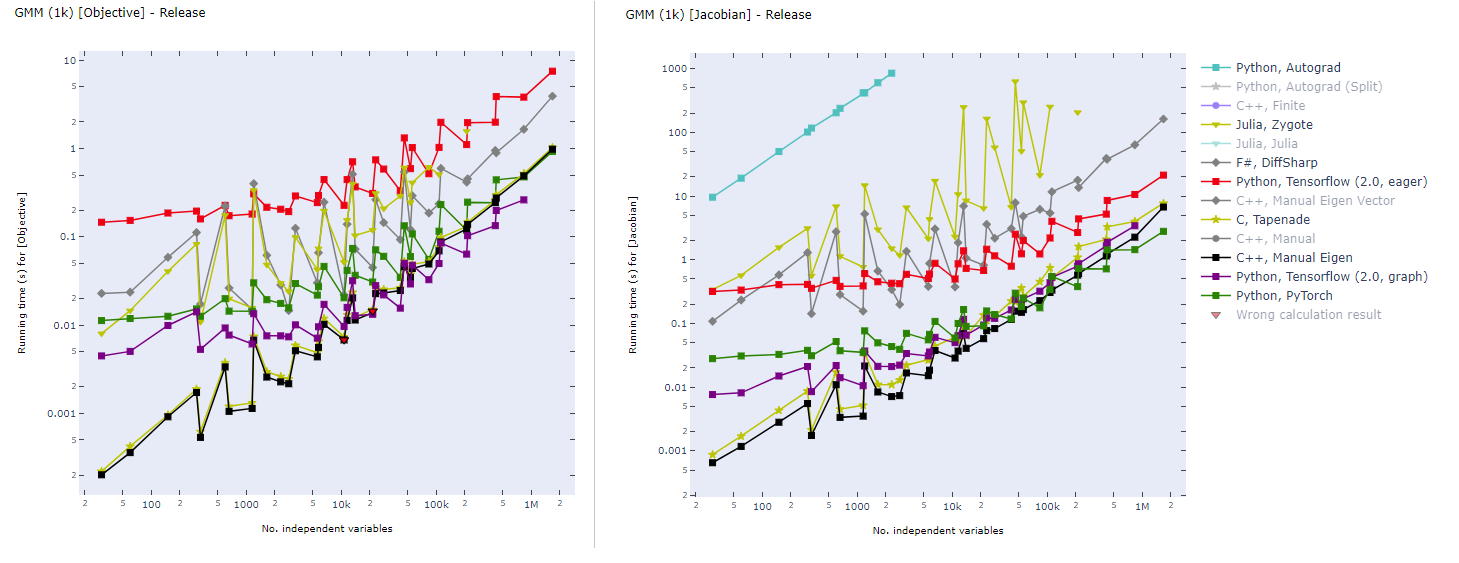
Static computational graph (graph-mode AD) enables compilation time optimization. Below is a benchmark of common AD software from here. In inverse modeling, we usually have a scalar-valued objective function, so the left panel is most relevant for ADCME.
Installation
- Install Julia.
🎉 Support Matrix
| | Julia≧1.3 | GPU | Custom Operator | |---------| ----- |-----|-----------------| | Linux |✔ | ✔ | ✔ | | MacOS |✔ | ✕ | ✔ | | Windows | ✔ | ✔ | ✔ |
- Install
ADCME
using Pkg
Pkg.add("ADCME")
❗ FOR WINDOWS USERS: See the instruction or the video for installation details.
❗ FOR MACOS USERS: See this troubleshooting list for potential installation and compilation problems on Mac.
- (Optional) Test
ADCME.jl
using Pkg
Pkg.test("ADCME")
See Troubleshooting if you encounter any compilation problems.
- (Optional) To enable GPU support, make sure
nvccis available from your environment (e.g., typenvccin your shell and you should get the location of the executable binary file), and then type
ENV["GPU"] = 1
Pkg.build("ADCME")
You can check the status with using ADCME; gpu_info().
- (Optional) Check the health of your installed ADCME and install missing dependencies or fixing incorrect paths.
using ADCME
doctor()
For manual installation without access to the internet, see here.
Install via Docker
If you have Docker installed on your system, you can try the no-hassle way to install ADCME (you don't even have to install Julia!):
docker run -ti kailaix/adcme
For GPU-enabled ADCME, use
docker run -ti --gpus all kailaix/adcme:gpu
See this guide for details. The docker images are hosted here.
Tutorial
Here we present three inverse problem examples. The first one is a parameter estimation problem, the second one is a function inverse problem where the unknown function does not depend on the state variables, and the third one is also a function inverse problem, but the unknown function depends on the state variables.
Parameter Inverse Problem
Consider solving the following problem
where
Assume that we have observed u(0.5)=1, we want to estimate b. In this case, he true value should be b=1.
using LinearAlgebra
using ADCME
n = 101 # number of grid nodes in [0,1]
h = 1/(n-1)
x = LinRange(0,1,n)[2:end-1]
b = Variable(10.0) # we use Variable keyword to mark the unknowns
A = diagm(0=>2/h^2*ones(n-2), -1=>-1/h^2*ones(n-3), 1=>-1/h^2*ones(n-3))
B = b*A + I # I stands for the identity matrix
f = @. 4*(2 + x - x^2)
u = B\f # solve the equation using built-in linear solver
ue = u[div(n+1,2)] # extract values at x=0.5
loss = (ue-1.0)^2
# Optimization
sess = Session(); init(sess)
BFGS!(sess, loss)
println("Estimated b = ", run(sess, b))
Expected output
Estimated b = 0.9995582304494237
The gradients can be obtained very easily. For example, if we want the gradients of loss with respect to b, the following code will create a Tensor for the gradient
julia> gradients(loss, b)
PyObject <tf.Tensor 'gradients_1/Mul_grad/Reshape:0' shape=() dtype=float64>
Function Inverse Problem: Full Field Data
Consider a nonlinear PDE,
where
Here f(x) can be computed from an analytical solution
In this problem, we are given the full field data of u(x) (the discrete value of u(x) is given on a very fine grid) and want to estimate the nonparametric function b(u). We approximate b(u) using a neural network and use the residual minimization method to find the optimal weig
Related Skills
YC-Killer
2.7kA library of enterprise-grade AI agents designed to democratize artificial intelligence and provide free, open-source alternatives to overvalued Y Combinator startups. If you are excited about democratizing AI access & AI agents, please star ⭐️ this repository and use the link in the readme to join our open source AI research team.
best-practices-researcher
The most comprehensive Claude Code skills registry | Web Search: https://skills-registry-web.vercel.app
research_rules
Research & Verification Rules Quote Verification Protocol Primary Task "Make sure that the quote is relevant to the chapter and so you we want to make sure that we want to have it identifie
groundhog
398Groundhog's primary purpose is to teach people how Cursor and all these other coding agents work under the hood. If you understand how these coding assistants work from first principles, then you can drive these tools harder (or perhaps make your own!).
