Goeffel
A tool for measuring the resource utilization of a specific process over time
Install / Use
/learn @jgehrcke/GoeffelREADME
Overview
Measures the resource utilization of a specific process over time.
Also measures the utilization/saturation of system-wide resources: this helps putting the process-specific metrics into context.
Built for Linux. Windows and Mac OS support might come.
For a list of the currently supported metrics see below.
Highlights
- Comes with a data plotting tool separate from the data acquisition program.
- High sampling rate: the default sampling interval of
0.5 smakes narrow spikes visible. - Values measurement correctness highly (see technical notes).
- Values interoperability: data files can be read with any HDF5 reader such as with pandas.read_hdf(), or with PyTables. See tips and tricks.
- Can monitor a program subject to process ID changes (for longevity experiments where the monitored process may occasionally restart).
- Can run indefinitely, with predictable disk space requirements (output file rotation and retention policy).
- Keeps your data organized: the time series data is written into a structured HDF5 file annotated with metadata (including program invocation time, system hostname, a custom label, the Goeffel software version, and others).
The name, Göffel, is German for spork:

Convenient, right?
Download & installation
The latest release can be downloaded and installed from PyPI, via pip:
$ pip install goeffel
pip can also install the latest development version:
$ pip install git+https://github.com/jgehrcke/goeffel
CLI tutorial
goeffel: data acquisition
You can invoke Goeffel with the --pid <pid> argument.
In this mode, goeffel stops the measurement and terminates itself once the process with the given ID goes away. Example:
$ goeffel --pid 29019
[... snip ...]
190809-15:46:57.914 INFO: Updated HDF5 file: wrote 20 sample(s) in 0.01805 s
[... snip ...]
190809-15:56:13.842 INFO: Cannot inspect process: process no longer exists (pid=29019)
190809-15:56:13.843 INFO: Wait for producer buffer to become empty
190809-15:56:13.843 INFO: Wait for consumer process to terminate
190809-15:56:13.854 INFO: Updated HDF5 file: wrote 13 sample(s) in 0.01077 s
190809-15:56:13.856 INFO: Sample consumer process terminated
For measuring beyond the process lifetime use --pid-command <command>.
In the following example, I use the pgrep utility for discovering a certain process (which is based on stress in this case):
$ goeffel --pid-command 'pgrep stress --newest'
[... snip ...]
190809-15:47:47.337 INFO: New process ID from PID command: 25890
[... snip ...]
190809-15:47:57.863 INFO: Updated HDF5 file: wrote 20 sample(s) in 0.01805 s
190809-15:48:06.850 INFO: Cannot inspect process: process no longer exists (pid=25890)
190809-15:48:06.859 INFO: PID command returned non-zero
[... snip ...]
190809-15:48:09.916 INFO: PID command returned non-zero
190809-15:48:10.926 INFO: New process ID from PID command: 28086
190809-15:48:12.438 INFO: Updated HDF5 file: wrote 20 sample(s) in 0.01013 s
190809-15:48:22.446 INFO: Updated HDF5 file: wrote 20 sample(s) in 0.01062 s
[... snip ...]
In this mode, goeffel runs forever until manually terminated via SIGINT or SIGTERM.
Process ID changes are detected by periodically running the discovery command until it returns a valid process ID on stdout.
This is useful for longevity experiments where the monitored process occasionally restarts, for instance as of fail-over scenarios.
goeffel-analysis: data inspection and visualization
Note: goeffel-analysis provides an opinionated and limited approach to visualizing data. For advanced and thorough data analysis I recommend building a custom (maybe even ad-hoc) data analysis pipeline using pandas and matplotlib, or using the tooling of your choice.
Also note: The command line interface provided by goeffel-analysis,
especially for the plot commands, might change in the future. Suggestions for
improvement are welcome, of course.
goeffel-analysis inspect:
Use goeffel-analysis inspect <path-to-HDF5-file> for inspecting the contents
of a Goeffel output file. Example:
$ goeffel-analysis inspect mwst18-master1-journal_20190801_111952.hdf5
Measurement metadata:
System hostname: int-master1-mwt18.foo.bar
Invocation time (local): 20190801_111952
PID command: pgrep systemd-journal
PID: None
Sampling interval: 1.0 s
Table properties:
Number of rows: 24981
Number of columns: 38
Number of data points (rows*columns): 9.49E+05
First row's (local) time: 2019-08-01T11:19:53.613377
Last row's (local) time: 2019-08-01T18:52:49.954582
Time span: 7h 32m 56s
Column names:
unixtime
... snip ...
system_mem_inactive
goeffel-analysis plot: quickly plot data from a single time series file
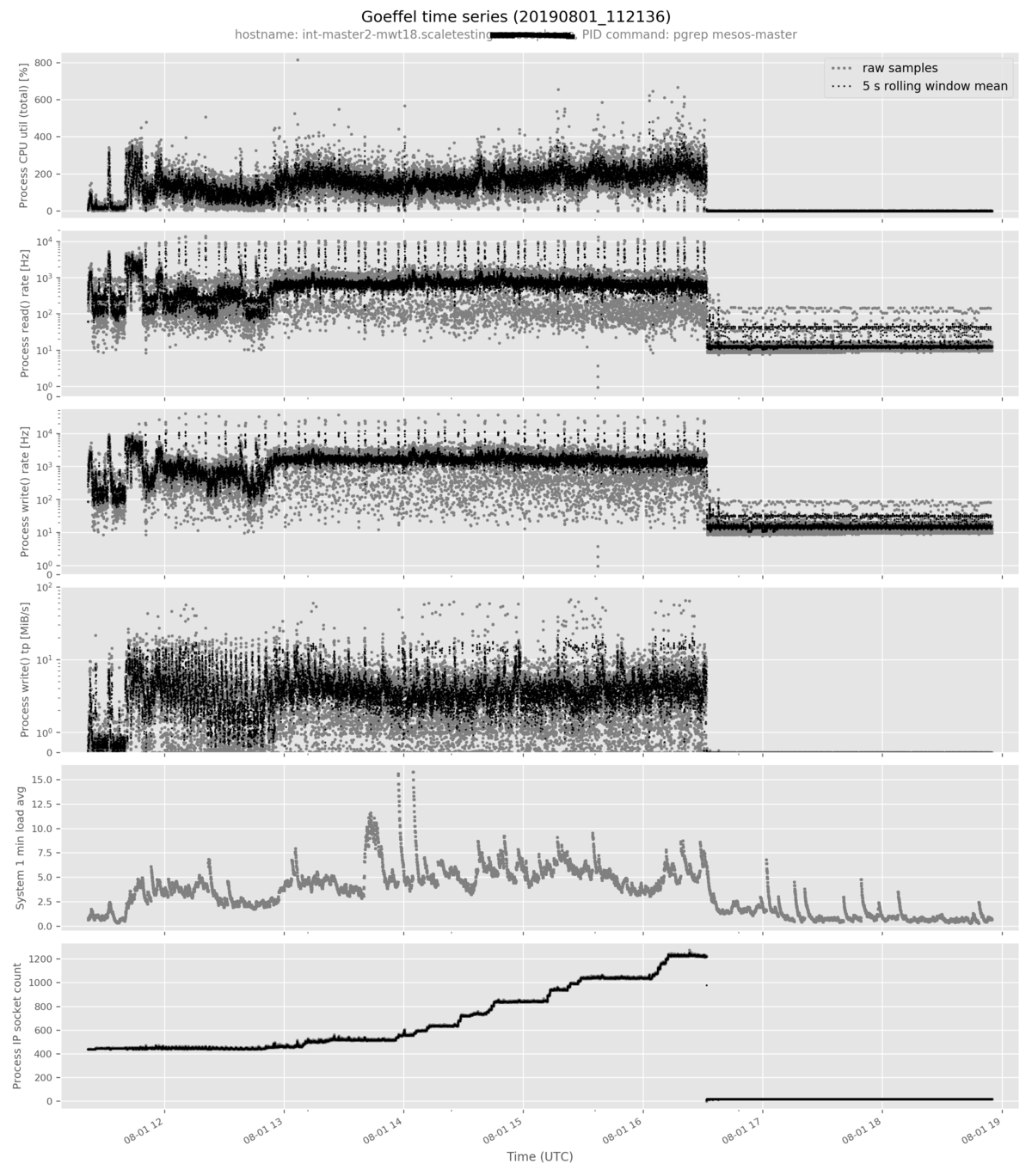
The goeffel-analysis plot <path-to-hdf5-file> command plots a pre-selected
set of metrics in an opinionated way. More metrics can be added to the plot with
the --metric <metric-name> option. Example command:
goeffel-analysis plot \
mwst18-master2-mesosmaster_20190801_112136.hdf5 \
--metric proc_num_ip_sockets_open
Example output figure:
goeffel-analysis flexplot: generic plot command
This command can be used for example for comparing multiple time series.
Say you have monitored the same program across multiple replicas in a distributed system and would like to compare the time evolution of a certain metric across these replicas.
Then the goeffel-analysis flexplot command is here to help, invoked with multiple --series arguments:
$ goeffel-analysis flexplot \
--series mwst18-master1-journal_20190801_111952.hdf5 master1 \
--series mwst18-master2-journal_20190801_112136.hdf5 master2 \
--series mwst18-master3-journal_20190801_112141.hdf5 master3 \
--series mwst18-master4-journal_20190801_112151.hdf5 master4 \
--series mwst18-master5-journal_20190801_112157.hdf5 master5 \
--column proc_cpu_util_percent_total \
'CPU util (total) / %' \
'systemd journal CPU utilization ' 15 \
--subtitle 'MWST18, measured with Goeffel' \
--legend-loc 'upper center'
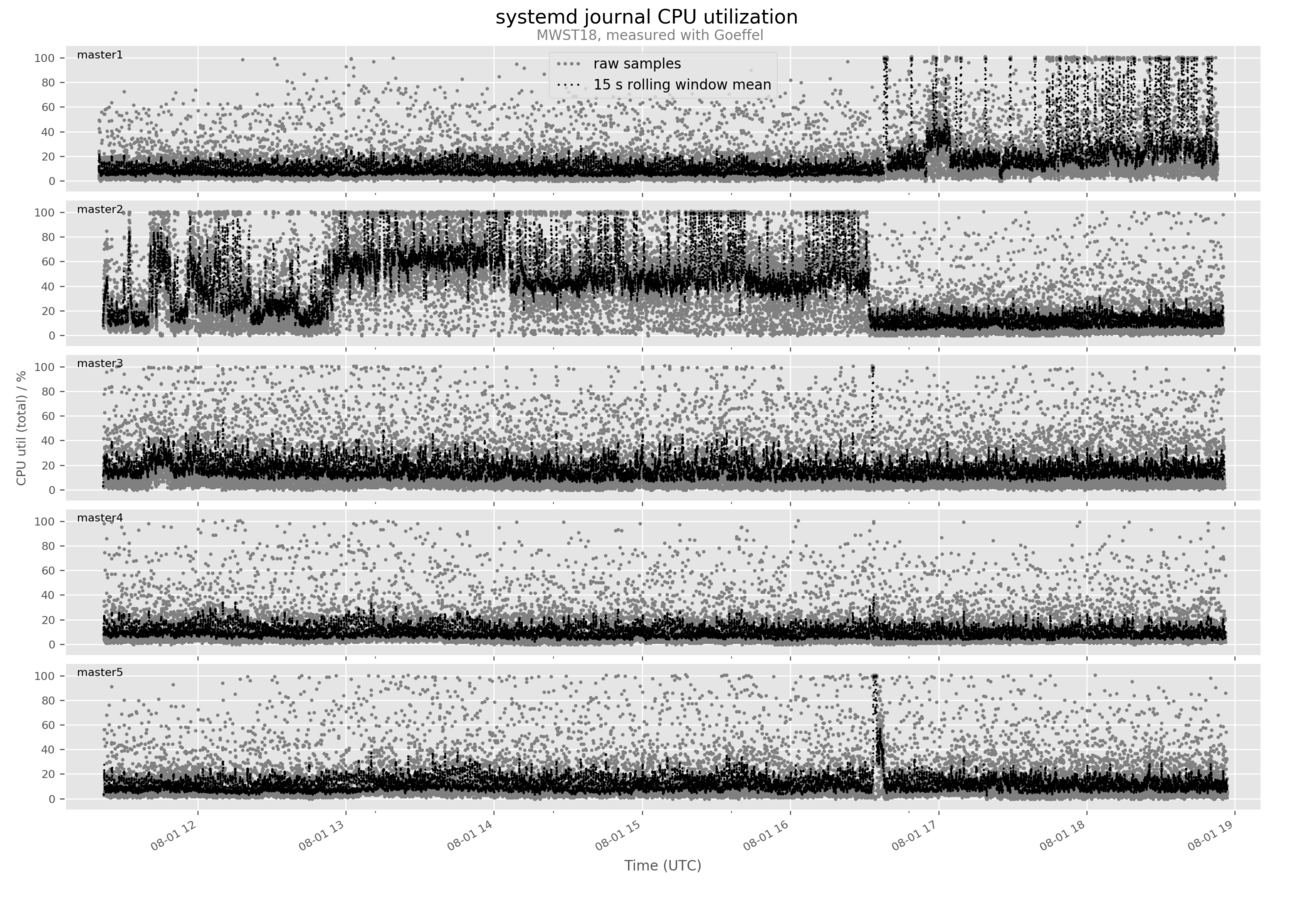
Example output figure:
Background and details
Prior art
This was born out of a need for solid tooling. We started with pidstat from
sysstat, launched as
pidstat -hud -p $PID 1 1. We found that it does not properly account for
multiple threads running in the same process and that various issues in that
regard exist in this program across various versions (see
here,
here, and
here).
The program cpustat open-sourced by Uber has a delightful README about the general measurement methodology and overall seems to be a great tool. However, it seems to be optimized for interactive usage (whereas we were looking for a robust measurement program which can be pointed at a process and then be left unattended for a significant while) and there does not seem to be a well-documented approach towards persisting the collected time series data on disk for later inspection.
The program psrecord (which effectively wraps psutil) has a similar fundamental approach as Goeffel; it however only measures few metrics, and it does not have a clear separation of concerns between persisting the data to disk, performing the measurement itself, and analyzing/plotting the data.
Technical notes
-
The core sampling loop does little work besides the measurement itself: it writes each sample to a queue. A separate process consumes this queue and persists the time series data to disk, for later inspection. This keeps the sampling rate predictable upon disk write latency spikes, or generally upon backpressure. This matters especially in cloud environments where we sometimes see fsync latencies of multiple seconds.
-
The sampling loop is (supposed to be, feedback welcome) built so that timing-related systematic measurement errors are minimized.
-
Goeffel tries to not asymmetrically hide measurement uncertainty. For example, you might see it measure a CPU utilization of a single-threaded process slightly larger than 100 %. That's simply the measurement error. In related tooling such as
sysstatit seems to be common practice to asymmetrically hide measurement uncertainty by capping values when they are known to in theory not exceed a certain threshold (example). -
goeffelmust be run withrootprivileges. -
The value
-1has a special meaning for some metrics (NaN, which cannot be represented properly in HDF5). Example: A disk write latency of-1 msmeans that no write happened in the corresponding time interval. -
Th
