Flashdeconv
Fast spatial deconvolution via leverage-score sketching — scales to million-spot datasets while preserving rare cell type signals.
Install / Use
/learn @cafferychen777/FlashdeconvREADME
FlashDeconv
Spatial deconvolution with linear scalability for atlas-scale data.
FlashDeconv estimates cell type proportions from spatial transcriptomics data (Visium, Visium HD, Stereo-seq). It is designed for large-scale analyses where computational efficiency is essential, while maintaining attention to low-abundance cell populations through leverage-score-based feature weighting.
Paper: Yang, C., Zhang, X. & Chen, J. FlashDeconv enables atlas-scale, multi-resolution spatial deconvolution via structure-preserving sketching. bioRxiv (2025). DOI: 10.64898/2025.12.22.696108
Installation
pip install flashdeconv
For development or additional I/O support, see Installation Options.
Quick Start
import scanpy as sc
import flashdeconv as fd
# Load data
adata_st = sc.read_h5ad("spatial.h5ad")
adata_ref = sc.read_h5ad("reference.h5ad")
# Deconvolve
fd.tl.deconvolve(adata_st, adata_ref, cell_type_key="cell_type")
# Results stored in adata_st.obsm["flashdeconv"]
sc.pl.spatial(adata_st, color="flashdeconv_Hepatocyte")
FlashDeconv is also available as a tool in ChatSpatial, an MCP server for spatial transcriptomics — run deconvolution through natural language from any compatible client.
Overview
Spatial deconvolution methods offer different trade-offs. Probabilistic approaches like Cell2Location and RCTD provide rigorous uncertainty quantification; methods like CARD incorporate spatial structure through dense kernel matrices. FlashDeconv takes a complementary approach, prioritizing computational efficiency for million-scale datasets.
Design Principles
-
Linear complexity — O(N) time and memory through randomized sketching and sparse graph regularization.
-
Leverage-based feature weighting — Variance-based selection (PCA, HVG) can underweight markers of low-abundance populations. We use leverage scores from the reference SVD to identify genes that define distinct transcriptomic directions, regardless of expression magnitude.
-
Sparse spatial regularization — Graph Laplacian smoothing with O(N) complexity, avoiding the O(N²) cost of dense kernel methods.
Performance
Scalability
| Spots | Time | Memory | |:------|:-----|:-------| | 10,000 | < 1 sec | < 1 GB | | 100,000 | ~4 sec | ~2 GB | | 1,000,000 | ~3 min | ~21 GB |
Benchmarked on MacBook Pro M2 Max (32GB unified memory), CPU-only.
Accuracy
On the Spotless benchmark:
| Metric | FlashDeconv | RCTD | Cell2Location | |:-------|:------------|:-----|:--------------| | Pearson (56 datasets) | 0.944 | 0.905 | 0.895 |
Performance varies by tissue type and experimental conditions. We recommend evaluating on data similar to your use case.
Algorithm
FlashDeconv solves a graph-regularized non-negative least squares problem:
minimize ½‖Y - βX‖²_F + ½λ·Tr(βᵀLβ) + ρ‖β‖₁, subject to β ≥ 0
where Y is spatial expression, X is reference signatures, L is the graph Laplacian, and β represents cell type abundances.
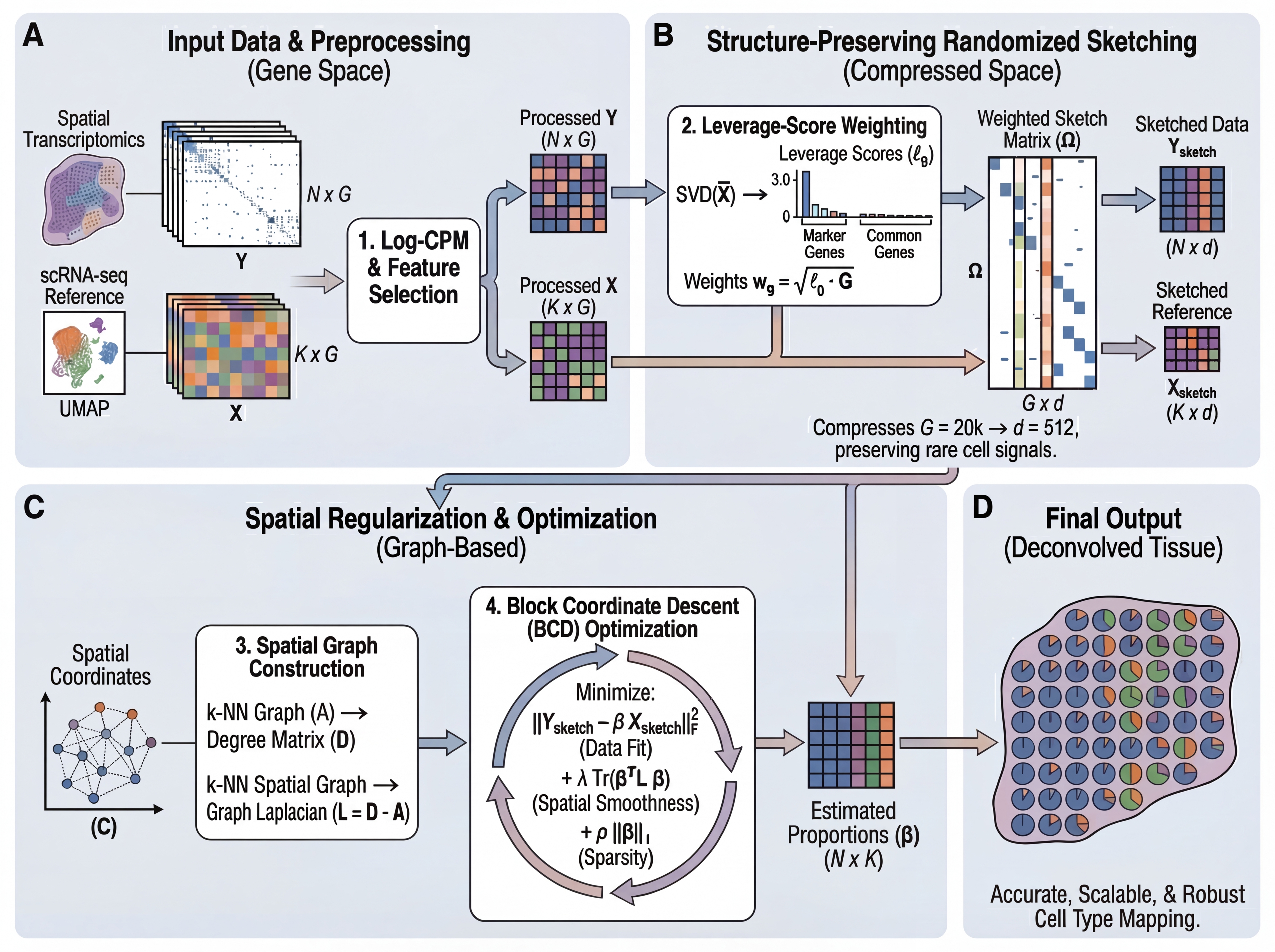
Pipeline:
- Select informative genes (HVG ∪ markers) and compute leverage scores
- Compress gene space via CountSketch with uniform hashing + leverage-weighted amplitudes (G → 512 dimensions)
- Construct sparse k-NN spatial graph
- Solve via block coordinate descent with spatial smoothing
API
Scanpy-style
fd.tl.deconvolve(
adata_st, # Spatial AnnData
adata_ref, # Reference AnnData
cell_type_key="cell_type", # Column in adata_ref.obs
key_added="flashdeconv", # Key for results
)
NumPy
from flashdeconv import FlashDeconv
model = FlashDeconv(
sketch_dim=512,
lambda_spatial="auto",
n_hvg=2000,
k_neighbors=6,
random_state=0,
)
proportions = model.fit_transform(Y, X, coords)
Parameters
| Parameter | Default | Description |
|:----------|:--------|:------------|
| sketch_dim | 512 | Sketch dimension |
| lambda_spatial | "auto" | Spatial regularization (auto-tuned) |
| n_hvg | 2000 | Highly variable genes |
| spatial_method | "knn" | Graph method: "knn", "radius", or "grid" |
| k_neighbors | 6 | Spatial graph neighbors (for "knn") |
| radius | None | Neighbor radius (required for "radius") |
| preprocess | "log_cpm" | Normalization: "log_cpm", "pearson", or "raw" |
| random_state | 0 | Random seed for reproducibility |
Output
| Attribute | Description |
|:----------|:------------|
| proportions_ | Cell type proportions (N × K), sum to 1 |
| beta_ | Raw abundances (N × K) |
| info_ | Convergence statistics |
Input Formats
- Spatial data: AnnData, NumPy array (N × G), or SciPy sparse matrix
- Reference: AnnData (aggregated by cell type) or NumPy array (K × G)
- Coordinates: Extracted from
adata.obsm["spatial"]or NumPy array (N × 2)
Reference Quality
Deconvolution accuracy depends on reference quality:
| Requirement | Guideline | |:------------|:----------| | Cells per type | ≥ 500 recommended | | Marker fold-change | ≥ 5× for distinguishability | | Signature correlation | < 0.95 between types | | No Unknown cells | Filter before deconvolution |
Critical: Always remove cells labeled "Unknown", "Unassigned", or similar. These cells act as universal signatures that absorb proportions from specific types—a fundamental property of regression-based deconvolution, not a FlashDeconv limitation.
See Reference Data Guide for details.
Installation Options
# Standard
pip install flashdeconv
# With AnnData support
pip install flashdeconv[io]
# Development
git clone https://github.com/cafferychen777/flashdeconv.git
cd flashdeconv && pip install -e ".[dev]"
Requirements: Python ≥ 3.9, numpy, scipy, numba. Optional: scanpy, anndata.
Citation
If you use FlashDeconv in your research, please cite:
Yang, C., Zhang, X. & Chen, J. FlashDeconv enables atlas-scale, multi-resolution spatial deconvolution via structure-preserving sketching. bioRxiv (2025). DOI: 10.64898/2025.12.22.696108
@article{yang2025flashdeconv,
title={FlashDeconv enables atlas-scale, multi-resolution spatial deconvolution
via structure-preserving sketching},
author={Yang, Chen and Zhang, Xianyang and Chen, Jun},
journal={bioRxiv},
year={2025},
doi={10.64898/2025.12.22.696108}
}
Resources
- Paper reproducibility code
- Reference data guide — Building quality reference signatures
- Stereo-seq guide — Platform-specific considerations
- GitHub Issues
- BSD-3-Clause License
Acknowledgments
We thank the developers of Spotless, Cell2Location, RCTD, CARD, and other deconvolution methods whose work contributed to this field.



