Locuscomparer
LocusCompareR is a R package with visualization tools for comparing two genetic association datasets.
Install / Use
/learn @boxiangliu/LocuscomparerREADME
LocusCompareR
1. Installation
LocusCompareR is an R package for visualization of GWAS-eQTL colocalization events.
Use the following commands to install LocusCompareR. If you don't have devtools, uncomment the first line to install it.
# install.packages("devtools")
devtools::install_github("boxiangliu/locuscomparer")
2. Example
To illustrate the use of locuscompare, we use the GWAS dataset from Nikpay et al. (2015) and the coronary artery eQTL dataset from GTEx v7 at the PHACTR1 locus:
library(locuscomparer)
gwas_fn = system.file('extdata','gwas.tsv', package = 'locuscomparer')
eqtl_fn = system.file('extdata','eqtl.tsv', package = 'locuscomparer')
locuscompare(in_fn1 = gwas_fn, in_fn2 = eqtl_fn, title1 = 'CAD GWAS', title2 = 'Coronary Artery eQTL')
The output from the main function is a figure like the following:
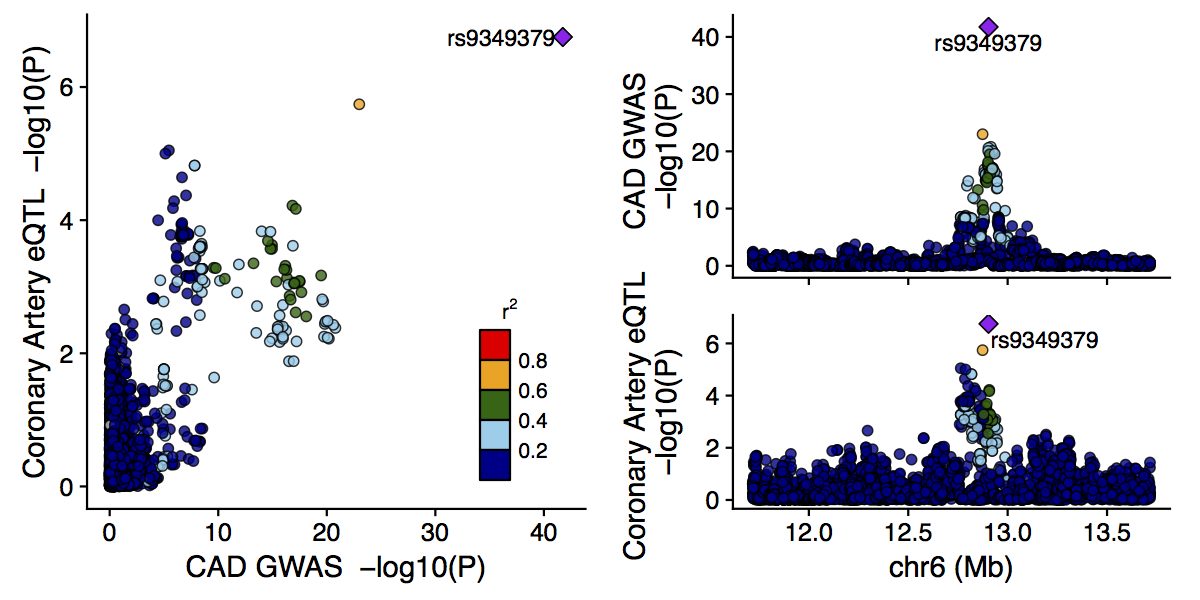
The labeled SNP is the lead SNP (in this case for both studies), and other SNPs are colored according to their LD $r^2$ with the lead SNP.
3. Using your own dataset:
The input to locuscompare::main() is a two-column tab-delimited text file with two columns:
- rsid
- pval
Here is an example file:
rsid pval
rs62156064 0.564395
rs7562234 0.399642
rs11677377 0.34308
rs35076156 0.625237
You can download the example files below: GWAS and eQTL datasets.
Then run the following commands:
library(locuscomparer)
gwas_fn = 'path/to/gwas.tsv'
eqtl_fn = 'path/to/eqtl.tsv'
locuscompare(in_fn1 = gwas_fn, in_fn2 = eqtl_fn, title = 'GWAS', title2 = 'eQTL')
4. Documentations
To view documentation for each function, type ?[function name] in the R console.
LocusCompareR current export the following functions:
Data munging
assign_color: Assign color to each SNP according to LD.get_lead_snp: Add a column of SNP labels to input data.frame.get_position: Append two columns, chromosome (chr) and position (pos), to the input data.frame.
Plotting
locuscompare: Make a locuscompare plot.make_combined_plot: Generated a combined plot with two locuszoom plots and a locuscompare plot.make_locuszoom: Make a simple locuszoom plot.make_scatterplot: Make a scatter plot (called the LocusCompare plot)
Data loading
read_metal" Read association summary statistics from file.retrieve_LD: Retrive SNP pairwise LD from database.
5. Citation
If you use locuscompare, please cite the following paper: https://www.nature.com/articles/s41588-019-0404-0
Boxiang Liu, Michael J. Gloudemans, Abhiram S. Rao, Erik Ingelsson & Stephen B. Montgomery (2019) Abundant associations with gene expression complicate GWAS follow-up, Nature Genetics
Related Skills
node-connect
342.5kDiagnose OpenClaw node connection and pairing failures for Android, iOS, and macOS companion apps
frontend-design
85.3kCreate distinctive, production-grade frontend interfaces with high design quality. Use this skill when the user asks to build web components, pages, or applications. Generates creative, polished code that avoids generic AI aesthetics.
openai-whisper-api
342.5kTranscribe audio via OpenAI Audio Transcriptions API (Whisper).
qqbot-media
342.5kQQBot 富媒体收发能力。使用 <qqmedia> 标签,系统根据文件扩展名自动识别类型(图片/语音/视频/文件)。
