NeuralGenetic
Building and training artificial neural networks (regression or classification) using the genetic algorithm.
Install / Use
/learn @ahmedfgad/NeuralGeneticREADME
NeuralGenetic: Training Neural Networks using the Genetic Algorithm
NeuralGenetic is a Python project for training neural networks using the genetic algorithm.
NeuralGenetic is part of the PyGAD library which is an open-source Python 3 library for implementing the genetic algorithm and optimizing machine learning algorithms. Both regression and classification neural networks are supported starting from PyGAD 2.7.0.
Check documentation of the NeuralGenetic project in the PyGAD's documentation: https://pygad.readthedocs.io/en/latest/gann.html
The library is under active development and more features are added regularly. If you want a feature to be supported, please check the Contact Us section to send a request.
Donation
- Credit/Debit Card: https://donate.stripe.com/eVa5kO866elKgM0144
- Open Collective: opencollective.com/pygad
- PayPal: Use either this link: paypal.me/ahmedfgad or the e-mail address ahmed.f.gad@gmail.com
- Interac e-Transfer: Use e-mail address ahmed.f.gad@gmail.com
Tutorial Project
IMPORTANT If you are coming for the code of the tutorial titled Artificial Neural Networks Optimization using Genetic Algorithm with Python, then it has been moved to the [Tutorial Project](https://github.com/ahmedfgad/NeuralGenetic/tree/master/Tutorial Project) directory on 15 May 2020.
Installation
To install PyGAD, simply use pip to download and install the library from PyPI (Python Package Index). The library is at PyPI at this page https://pypi.org/project/pygad.
Install PyGAD with the following command:
pip install pygad
To get started with PyGAD, please read the documentation at Read The Docs https://pygad.readthedocs.io.
PyGAD Source Code
The source code of the PyGAD' modules is found in the following GitHub projects:
- pygad: (https://github.com/ahmedfgad/GeneticAlgorithmPython)
- pygad.nn: https://github.com/ahmedfgad/NumPyANN
- pygad.gann: https://github.com/ahmedfgad/NeuralGenetic
- pygad.cnn: https://github.com/ahmedfgad/NumPyCNN
- pygad.gacnn: https://github.com/ahmedfgad/CNNGenetic
- pygad.kerasga: https://github.com/ahmedfgad/KerasGA
- pygad.torchga: https://github.com/ahmedfgad/TorchGA
The documentation of PyGAD is available at Read The Docs https://pygad.readthedocs.io.
PyGAD Documentation
The documentation of the PyGAD library is available at Read The Docs at this link: https://pygad.readthedocs.io. It discusses the modules supported by PyGAD, all its classes, methods, attribute, and functions. For each module, a number of examples are given.
If there is an issue using PyGAD, feel free to post at issue in this GitHub repository https://github.com/ahmedfgad/GeneticAlgorithmPython or by sending an e-mail to ahmed.f.gad@gmail.com.
If you built a project that uses PyGAD, then please drop an e-mail to ahmed.f.gad@gmail.com with the following information so that your project is included in the documentation.
- Project title
- Brief description
- Preferably, a link that directs the readers to your project
Please check the Contact Us section for more contact details.
Life Cycle of PyGAD
The next figure lists the different stages in the lifecycle of an instance of the pygad.GA class. Note that PyGAD stops when either all generations are completed or when the function passed to the on_generation parameter returns the string stop.
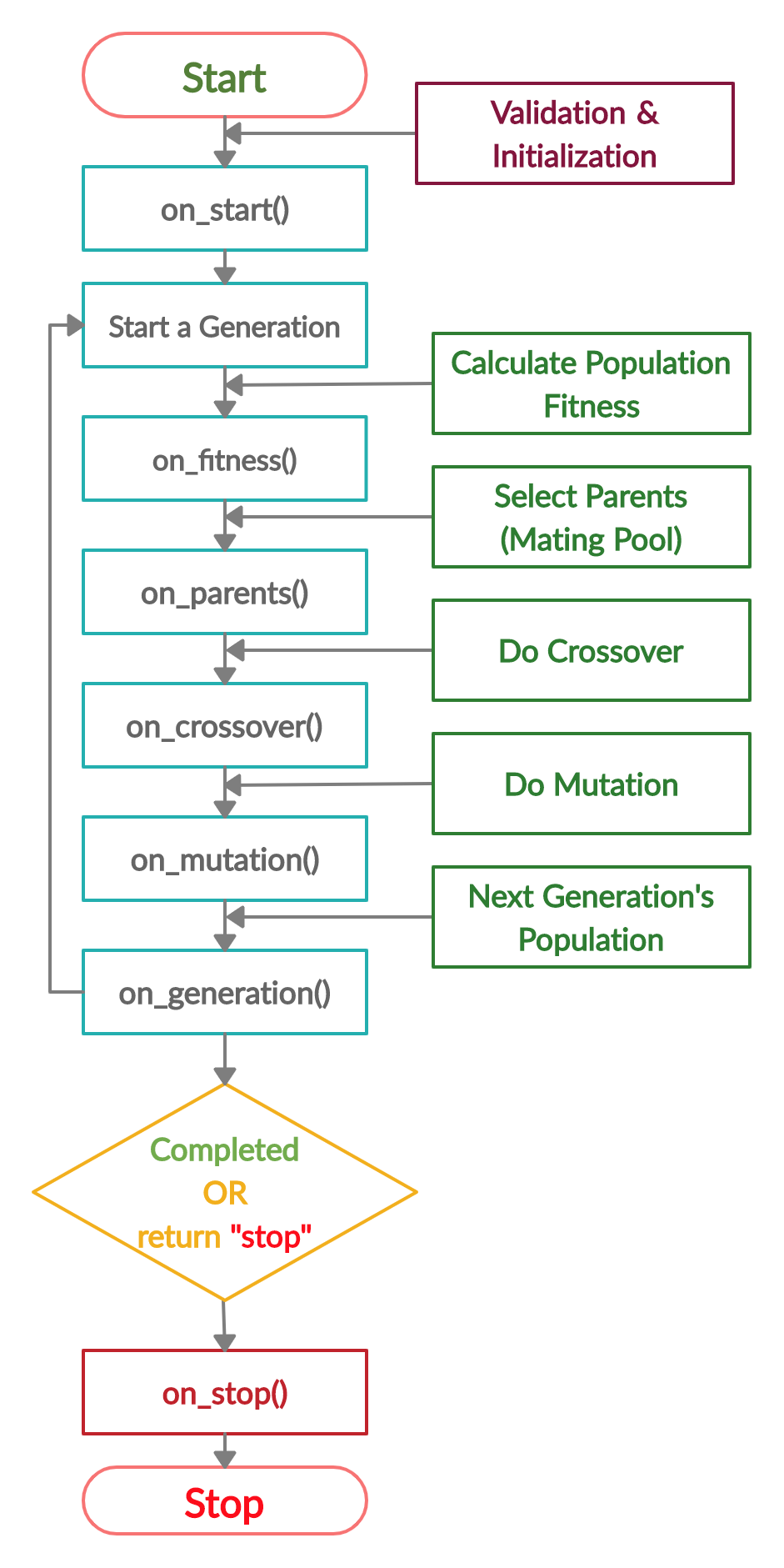
The next code implements all the callback functions to trace the execution of the genetic algorithm. Each callback function prints its name.
import pygad
import numpy
function_inputs = [4,-2,3.5,5,-11,-4.7]
desired_output = 44
def fitness_func(ga_instance, solution, solution_idx):
output = numpy.sum(solution*function_inputs)
fitness = 1.0 / (numpy.abs(output - desired_output) + 0.000001)
return fitness
fitness_function = fitness_func
def on_start(ga_instance):
print("on_start()")
def on_fitness(ga_instance, population_fitness):
print("on_fitness()")
def on_parents(ga_instance, selected_parents):
print("on_parents()")
def on_crossover(ga_instance, offspring_crossover):
print("on_crossover()")
def on_mutation(ga_instance, offspring_mutation):
print("on_mutation()")
def on_generation(ga_instance):
print("on_generation()")
def on_stop(ga_instance, last_population_fitness):
print("on_stop()")
ga_instance = pygad.GA(num_generations=3,
num_parents_mating=5,
fitness_func=fitness_function,
sol_per_pop=10,
num_genes=len(function_inputs),
on_start=on_start,
on_fitness=on_fitness,
on_parents=on_parents,
on_crossover=on_crossover,
on_mutation=on_mutation,
on_generation=on_generation,
on_stop=on_stop)
ga_instance.run()
Based on the used 3 generations as assigned to the num_generations argument, here is the output.
on_start()
on_fitness()
on_parents()
on_crossover()
on_mutation()
on_generation()
on_fitness()
on_parents()
on_crossover()
on_mutation()
on_generation()
on_fitness()
on_parents()
on_crossover()
on_mutation()
on_generation()
on_stop()
Example
Check the PyGAD's documentation for information about the implementation of this example.
import numpy
import pygad
import pygad.nn
import pygad.gann
def fitness_func(ga_instance, solution, sol_idx):
global GANN_instance, data_inputs, data_outputs
predictions = pygad.nn.predict(last_layer=GANN_instance.population_networks[sol_idx],
data_inputs=data_inputs)
correct_predictions = numpy.where(predictions == data_outputs)[0].size
solution_fitness = (correct_predictions/data_outputs.size)*100
return solution_fitness
def callback_generation(ga_instance):
global GANN_instance, last_fitness
population_matrices = pygad.gann.population_as_matrices(population_networks=GANN_instance.population_networks,
population_vectors=ga_instance.population)
GANN_instance.update_population_trained_weights(population_trained_weights=population_matrices)
print("Generation = {generation}".format(generation=ga_instance.generations_completed))
print("Fitness = {fitness}".format(fitness=ga_instance.best_solution()[1]))
print("Change = {change}".format(change=ga_instance.best_solution()[1] - last_fitness))
last_fitness = ga_instance.best_solution()[1].copy()
# Holds the fitness value of the previous generation.
last_fitness = 0
# Preparing the NumPy array of the inputs.
data_inputs = numpy.array([[1, 1],
[1, 0],
[0, 1],
[0, 0]])
# Preparing the NumPy array of the outputs.
data_outputs = numpy.array([0,
1,
1,
0])
# The length of the input vector for each sample (i.e. number of neurons in the input layer).
num_inputs = data_inputs.shape[1]
# The number of neurons in the output layer (i.e. number of classes).
num_classes = 2
# Creating an initial population of neural networks. The return of the initial_population() function holds references to the networks, not their weights. Using such references, the weights of all networks can be fetched.
num_solutions = 6 # A solution or a network can be used interchangeably.
GANN_instance = pygad.gann.GANN(num_solutions=num_solutions,
num_neurons_input=num_inputs,
num_neurons_hidden_layers=[2],
num_neurons_output=num_classes,
hidden_activations=["relu"],
output_activation="softmax")
# population does not hold the numerical weights of the network instead it holds a list of references to each last layer of each network (i.e. solution) in the population. A solution or a network can be used interchangeably.
# If there is a population with 3 solutions (i.e. networks), then the population is a list with 3 elements. Each element is a reference to the last layer of each network. Using such a reference, all details of the network can be accessed.
population_vectors = pygad.gann.population_as_vectors(population_networks=GANN_instance.population_networks)
# To prepare the initial population, there are 2 ways:
# 1) Prepare it yourself and pass it to the initial_population parameter. This way is useful when the user wants to start the genetic algorithm with a custom initial population.
# 2) Assign valid integer values to the sol_per_pop and num_genes parameters. If the initial_population parameter exists, then the sol_per_pop and num_genes parameters
