Methylartist
Tools for plotting methylation data in various ways
Install / Use
/learn @adamewing/MethylartistREADME
methylartist
Tools for parsing and plotting methylation patterns
Installation
Available through pip and conda:
pip install methylartist
or
conda install -c bioconda methylartist
Alternatively:
git clone https://github.com/adamewing/methylartist.git or download a .zip file from GitHub.
Various tests/examples are availble at methylartists-tests:
git clone https://github.com/adamewing/methylartist-tests.git
cd methylartist-tests
source run_tests.sh
Input data
Alignments (.bam)
Alignments stored in .bam format should be sorted and indexed and should use the same read names as the associated methylation data.
Modified Base Calls
The easiest way to provide modified basecall data is through .bam files with the MM and ML tags for modified base calling such as those produced by guppy or dorado. Note that the mod_mappings.bam file output by megalodon will work for modified base calling, but is unsuitable for downstream applications involving sequence variation, including phasing.
If .bam files with modified base calls are not available, methylartist has functions for loading methylation data can from nanopolish, megalodon, or basecalled guppy fast5s, using the appropriate function below. Assays that use C/U conversion for methylation inference are also supported through the db-sub command as noted below.
Once coverted, the sqlite .db file can be input to methylartist functions (e.g. segplot, locus, etc).
Commands:
db-nanopolish
Load nanopolish methylation into sqlite db.
Example:
Loading results from nanopolish call-methylation to a database:
methylartist db-nanopolish -m MCF7_ATCC_REP1.nanopolish.tsv.gz -d MCF7_ATCC.nanopolish.db
Appending additional results to the above database:
methylartist db-nanopolish -m MCF7_ATCC_REP2.nanopolish.tsv.gz -d MCF7_ATCC.nanopolish.db -a
Loading results with the current recommended cutoffs for nanopolish (abs(llr) > 2.0, scale grouped CpGs):
methylartist db-nanopolish -m MCF7_ATCC_REP1.nanopolish.tsv.gz,MCF7_ATCC_REP2.nanopolish.tsv.gz -d MCF7_ATCC.nanopolish.db -t 2.0 -s
Inputs can be uncompressed or .gzipped.
db-megalodon
Load megalodon methylation into sqlite db.
The input file is the output of megalodon_extras per_read_text modified_bases /path/to/megalodon_output, which needs to be run prior to this script.
The default filename (/path/to/megalodon_output/per_read_modified_base_calls.txt) is the same for all megalodon runs, so the --db option is recommended to make the output database more identifiable for downstream analysis.
Example:
methylartist db-megalodon -m MCF7_ATCC_REP1/per_read_modified_base_calls.txt --db MCF7_ATCC.megalodon.db
Appending (-a) additional results to the above database:
methylartist db-megalodon -m MCF7_ATCC_REP2/per_read_modified_base_calls.txt --db MCF7_ATCC.megalodon.db -a
Input files can be uncompressed or .gzipped.
db-custom
This enables free-form parsing of modified basecall tables into methylartist .db files for tools where modified base .bam files are not available and certain requirements are met. The table must contain, at a minimum, the read names, genomic position (chromosome and position), strand, and probability of the target base being modified. If not specified by a column (--modbasecol), the modification is specified by --modbase. The probability is assumed to be a raw probability between 0 and 1 of a given base being modified i.e. 1-p(canonical), other schemes may be used but --canprob and --mincanprob must be specified to set a column for canoncial base scores and a cutoff for a base being canonical.
For example, modified basecalls from deepsignal-plant can be loaded as follows:
/home/taewing/methylartist/methylartist db-custom -m deepsignal_example.C.call_mods.tsv --readname 4 --chrom 0 --pos 1 --strand 2 --modprob 7 --modbase m -d deepsignal_example.db
db-sub
methylartist now supports C/T data if the .bam file is noted as being C/T substitution data via --ctbam (works for locus, region, segmeth, wgmeth)
As of 1.3.0, methylartist supports display of C/U base substitution data (i.e. WGBS or EM-seq data) via creation of a methylartist .db file, simply pass the .bam file as input and specify an output file:
methylartist db-sub -b NA12878.EMSEQ.GAPDH.bam -d NA12878.EMSEQ.GAPDH.db
Note that the .bam file has to include the MD tag (aligners for bisulfite/em-seq data should do this).
segmeth
Outputs aggregate methylation / demethylation call counts over intervals. Required before generating strip / violin plots with segplot
Requires whitespace-delimited list of segments in a BED3+2 format: chromosome, start, end, label, strand.
One or more .bam files may be supplied via the -b/--bams parameter. Multiple .bams may be comma-delimited.
Optional sample input file -d/--data has the following whitespace-delimited fields (one sample per line): BAM file, Methylation DB (generated with e.g. db-nanopolish)
Highly recommend parallelising with -p/--proc option if possible.
Can be used to generate genome-wide methylation stats aggregated over windows via bedtools makewindows.
Example:
Aggregate whole-genome CpG methylation in 10kbp bins, promoters (Eukaryotic Promoter Database, EPD), L1HS and SVA retrotransposons:
methylartist segmeth -d MCF7_data_megalodon.txt -i MCF7_megalodon_annotations.bed -p 32
Contents of MCF7_data_megalodon.txt:
MCF7_ATCC.haplotag.bam MCF7_ATCC.megalodon.db
MCF7_ECACC.haplotag.bam MCF7_ECACC.megalodon.db
Contents of MCF7_megalodon_annotations.bed (first 10 lines):
chr1 0 10000 WG_10kbp
chr1 10000 20000 WG_10kbp
chr1 20000 30000 WG_10kbp
chr1 30000 40000 WG_10kbp
chr1 40000 50000 WG_10kbp
chr1 50000 60000 WG_10kbp
chr1 60000 70000 WG_10kbp
chr1 70000 80000 WG_10kbp
chr1 80000 90000 WG_10kbp
chr1 90000 100000 WG_10kbp
Output in MCF7_megalodon_annotations.segmeth.tsv (first 10 lines):
seg_id seg_chrom seg_start seg_end seg_name seg_strand MCF7_ATCC.haplotag_m_meth_calls MCF7_ATCC.haplotag_m_unmeth_calls MCF7_ATCC.haplotag_m_no_calls MCF7_ATCC.haplotag_m_methfrac MCF7_ECACC.haplotag_m_meth_calls MCF7_ECACC.haplotag_m_unmeth_calls MCF7_ECACC.haplotag_m_no_calls MCF7_ECACC.haplotag_m_methfrac
chr1:0-10000 chr1 0 10000 WG_10kbp . 0 0 0 NaN 0 0 0 NaN
chr1:10000-20000 chr1 10000 20000 WG_10kbp . 4836 1205 893 0.8005297136235723 5994 1629 1254 0.7863046044864227
chr1:20000-30000 chr1 20000 30000 WG_10kbp . 1923 2641 802 0.42134092900964065 2093 3216 1032 0.39423620267470333
chr1:30000-40000 chr1 30000 40000 WG_10kbp . 974 790 273 0.5521541950113379 1331 821 416 0.6184944237918215
chr1:40000-50000 chr1 40000 50000 WG_10kbp . 361 398 149 0.4756258234519104 579 664 255 0.4658085277554304
chr1:50000-60000 chr1 50000 60000 WG_10kbp . 631 300 133 0.677765843179377 1086 472 242 0.6970474967907574
chr1:60000-70000 chr1 60000 70000 WG_10kbp . 315 494 130 0.38936959208899874 571 671 255 0.45974235104669886
chr1:70000-80000 chr1 70000 80000 WG_10kbp . 196 150 31 0.5664739884393064 288 214 79 0.5737051792828686
chr1:80000-90000 chr1 80000 90000 WG_10kbp . 297 122 29 0.7088305489260143 127 57 16 0.6902173913043478
segplot
Generates strip plots or violin plots (-v/--violin) from segmeth output.
Examples:
Strip plot of whole-genome CpG methylation in 10kbp bins, promoters (Eukaryotic Promoter Database, EPD), L1HS and SVA retrotransposons:
methylartist segplot -s MCF7_megalodon_annotations.segmeth.tsv

As above, but use violin plots:
methylartist segplot -s MCF7_megalogon_annotations.segmeth.tsv -v
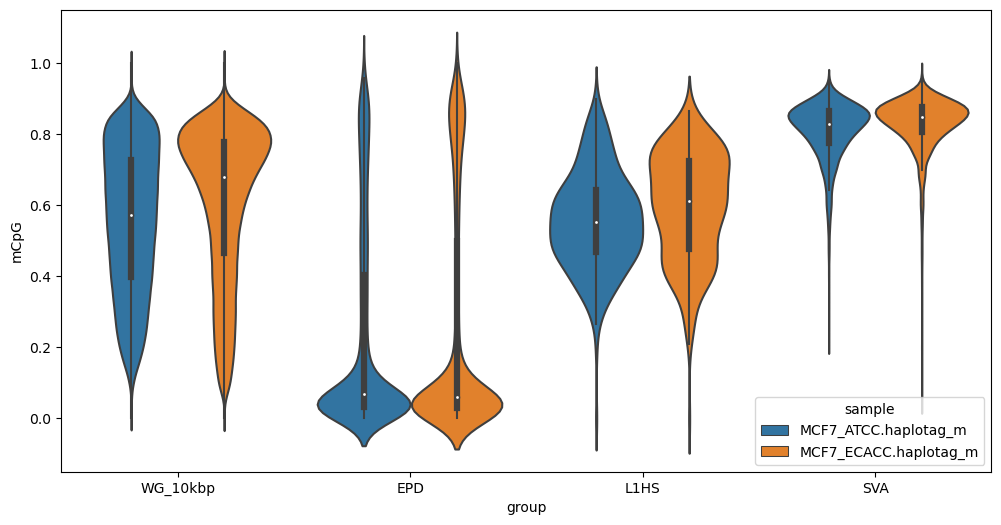
Note that default output is in .png format. For .svg vector output suitable for editing in inkscape or illustrator add the --svg option. Note that for strip plots, this is often inadvisable due to the large number of points.
New in 1.0.7, ridge plots (-g/--ridge):
methylartist segplot -s L1_FL.MCF7_data_megalodon.segmeth.tsv -c L1HS,L1PA2,L1PA3,L1PA4,L1PA5,L1PA6,L1PA7,L1PA8 -g --palette magma
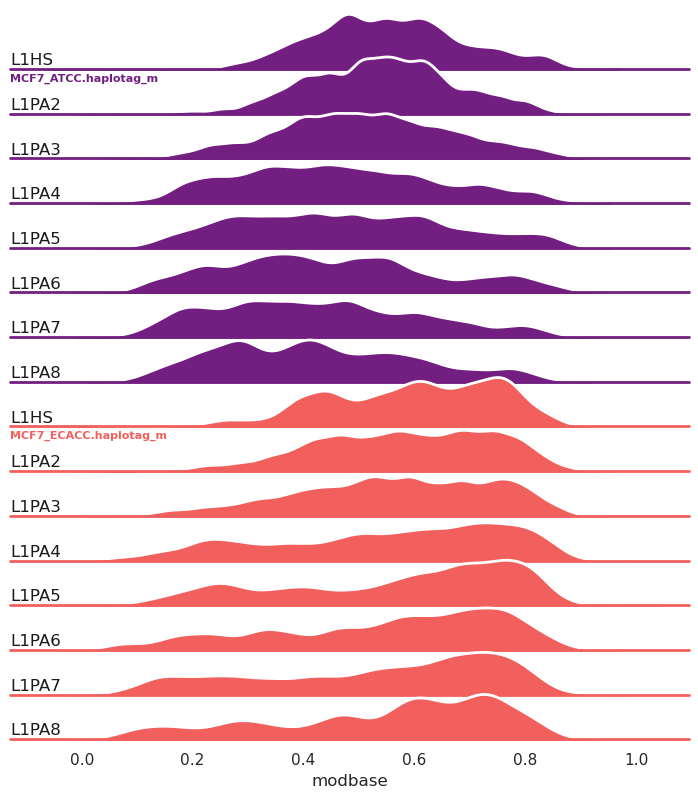
Ridge plots can also be grouped by annotation (--ridge_group_by_annotation) rather than by sample as in the above example.
locus
Generates smoothed methylation profiles across specific loci with many configurable parameters for one or more samples.
One or more .bam files (with Mm/Ml tags) may be supplied via the -b/--bams parameter. Multiple .bams may be comma-delimited.
Optional sample input file -d/--data has the following whitespace-delimited fields (one sample per line): BAM file, Methylation DB (generated with e.g. db-nanopolish)
Example:
Plot of the GPER1 locus in hg38, highlighting the GeneHancer promoter/enhancer annotation (GH07J001085).
methylartist locus -d MCF7_data_megalodon.txt -i chr7:1072064-1101499 -g Homo_sapiens.GRCh38
Related Skills
node-connect
341.8kDiagnose OpenClaw node connection and pairing failures for Android, iOS, and macOS companion apps
frontend-design
84.6kCreate distinctive, production-grade frontend interfaces with high design quality. Use this skill when the user asks to build web components, pages, or applications. Generates creative, polished code that avoids generic AI aesthetics.
openai-whisper-api
341.8kTranscribe audio via OpenAI Audio Transcriptions API (Whisper).
commit-push-pr
84.6kCommit, push, and open a PR
