ScMGCA
"Topological Identification and Interpretation for Single-cell Gene Regulation Elucidation across Multiple Platforms using scMGCA" in Nature Communications 2023
Install / Use
/learn @Philyzh8/ScMGCAREADME
scMGCA
scMGCA is a Python package containing tools for clustering single-cell data based on a graph-embedding autoencoder that simultaneously learns cell–cell topology representation and cluster assignments.
Overview
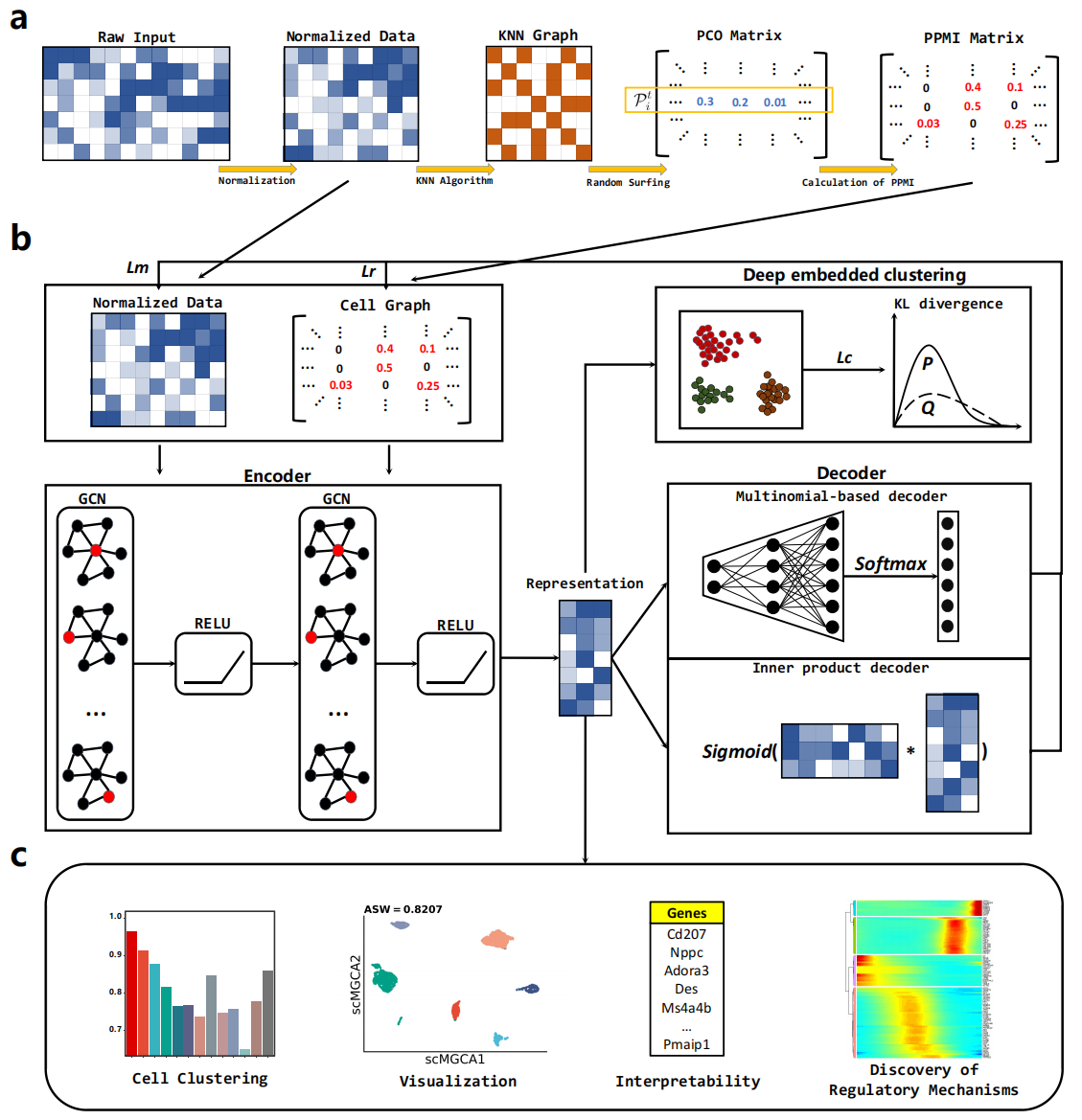
Single-cell RNA sequencing provides high-throughput gene expression information to explore cellular heterogeneity at the individual cell level. A major challenge in characterizing high-throughput gene expression data arises from challenges related to dimensionality, and the prevalence of dropout events. To address these concerns, we develop a deep graph learning method, scMGCA, for single-cell data analysis. scMGCA is based on a graph-embedding autoencoder that simultaneously learns cell-cell topology representation and cluster assignments. We show that scMGCA is accurate and effective for cell segregation and batch effect correction, outperforming other state-of-the-art models across multiple platforms. In addition, we perform genomic interpretation on the key compressed transcriptomic space of the graph-embedding autoencoder to demonstrate the underlying gene regulation mechanism. We demonstrate that in a pancreatic ductal adenocarcinoma dataset, scMGCA successfully provides annotations on the specific cell types and reveals differential gene expression levels across multiple tumor-associated and cell signalling pathways.
System Requirements
Hardware requirements
scMGCA package requires only a standard computer with enough RAM to support the in-memory operations.
Software requirements
OS Requirements
This package is supported for Linux. The package has been tested on the following systems:
- Linux: Ubuntu 18.04
Python Dependencies
scMGCA mainly depends on the Python scientific stack.
numpy
scipy
tensorflow
scikit-learn
pandas
scanpy
anndata
For specific setting, please see <a href="https://github.com/Philyzh8/scMGCA/blob/master/requirements.txt">requirement</a>.
Installation Guide:
Install from PyPi
$ conda create -n scMGCA_env python=3.6.8
$ conda activate scMGCA_env
$ pip install -r requirements.txt
$ pip install scMGCA
Usage
scMGCA is a deep graph embedding learning method for single-cell clustering, which can be used to:
- Single-cell data clustering. The example can be seen in the <a href="https://github.com/Philyzh8/scMGCA/blob/master/tutorial/demo.py">demo.py</a>.
- Correct the batch effect of data from different scRNA-seq protocols. The example can be seen in the <a href="https://github.com/Philyzh8/scMGCA/blob/master/tutorial/demo_batch.py">demo_batch.py</a>.
- Analysis of the mouse brain data with 1.3 million cells. The example can be seen in the <a href="https://github.com/Philyzh8/scMGCA/blob/master/tutorial/demo_scale.py">demo_scale.py</a>.
- Provide an automatic hyperparameter search algorithm. The example can be seen in the <a href="https://github.com/Philyzh8/scMGCA/blob/master/tutorial/demo_para.py">demo_para.py</a>.
We give users some suggestions for running in the <a href="https://github.com/Philyzh8/scMGCA/blob/master/tutorial/tutorial.md">tutorial.md</a>.
Data Availability
The real data sets we used can be download in <a href="https://doi.org/10.5281/zenodo.7475687">data</a>.
License
This project is covered under the MIT License.
Citation
@article{yu2023topological,
title={Topological identification and interpretation for single-cell gene regulation elucidation across multiple platforms using scMGCA},
author={Yu, Zhuohan and Su, Yanchi and Lu, Yifu and Yang, Yuning and Wang, Fuzhou and Zhang, Shixiong and Chang, Yi and Wong, Ka-Chun and Li, Xiangtao},
journal={Nature Communications},
volume={14},
number={1},
pages={400},
year={2023},
publisher={Nature Publishing Group UK London}
}
Related Skills
node-connect
338.0kDiagnose OpenClaw node connection and pairing failures for Android, iOS, and macOS companion apps
frontend-design
83.4kCreate distinctive, production-grade frontend interfaces with high design quality. Use this skill when the user asks to build web components, pages, or applications. Generates creative, polished code that avoids generic AI aesthetics.
openai-whisper-api
338.0kTranscribe audio via OpenAI Audio Transcriptions API (Whisper).
commit-push-pr
83.4kCommit, push, and open a PR


