Fastplong
Ultra-fast preprocessing and quality control for long-read sequencing data
Install / Use
/learn @OpenGene/FastplongREADME
fastplong
Ultrafast preprocessing and quality control for long reads (Nanopore, PacBio, Cyclone, etc.).
If you're searching for tools to preprocess short reads (Illumina, MGI, etc.), please use fastp
fastplong supports batch processing of multiple FASTQ files in a folder, see - batch processing
- simple usage
- examples of report
- get fastplong
- input and output
- fastplong workflow
- filtering
- adapters
- per read cutting by quality score
- break read to subreads by discarding low quality regions
- mask low quality regions with N
- global trimming
- output splitting
- batch processing
- all options
simple usage
fastplong -i in.fq -o out.fq
Both input and output can be gzip compressed. By default, the HTML report is saved to fastplong.html (can be specified with -h option), and the JSON report is saved to fastplong.json (can be specified with -j option).
examples of report
fastplong creates reports in both HTML and JSON format.
- HTML report: http://opengene.org/fastplong/fastplong.html
- JSON report: http://opengene.org/fastplong/fastplong.json
get fastplong
install with Bioconda
conda install -c bioconda fastplong
download the latest prebuilt binary for Linux users
This binary was compiled on CentOS, and tested on CentOS/Ubuntu
# download the latest build
wget http://opengene.org/fastplong/fastplong
chmod a+x ./fastplong
# or download specified version, i.e. fastplong v0.2.2
wget http://opengene.org/fastplong/fastplong.0.2.2
mv fastplong.0.2.2 fastplong
chmod a+x ./fastplong
or compile from source
fastplong depends on libdeflate and isa-l for fast decompression and compression of zipped data, and depends on libhwy for SIMD acceleration. It's recommended to install all of them via Anaconda:
conda install conda-forge::libdeflate
conda install conda-forge::isa-l
conda install conda-forge::libhwy
You can also try to install them with other package management systems like apt/yum on Linux, or brew on MacOS. Otherwise you can compile them from source (https://github.com/intel/isa-l, https://github.com/ebiggers/libdeflate, and https://github.com/google/highway)
download and build fastplong
# get source (you can also use browser to download from master or releases)
git clone https://github.com/OpenGene/fastplong.git
# build
cd fastplong
make -j
# test
make test
# Install
sudo make install
input and output
Specify input by -i or --in, and specify output by -o or --out.
- if you don't specify the output file names, no output files will be written, but the QC will still be done for both data before and after filtering.
- the output will be gzip-compressed if its file name ends with
.gz
output to STDOUT
fastplong supports streaming the passing-filter reads to STDOUT, so that it can be passed to other compressors like bzip2, or be passed to aligners like minimap2 or bowtie2.
- specify
--stdoutto enable this mode to stream output to STDOUT
input from STDIN
- specify
--stdinif you want to read the STDIN for processing.
store the reads that fail the filters
- give
--failed_outto specify the file name to store the failed reads. - if one read failed and is written to
--failed_out, itsfailure reasonwill be appended to its read name. For example,failed_quality_filter,failed_too_shortetc.
process only part of the data
If you don't want to process all the data, you can specify --reads_to_process to limit the reads to be processed. This is useful if you want to have a fast preview of the data quality, or you want to create a subset of the filtered data.
do not overwrite exiting files
You can enable the option --dont_overwrite to protect the existing files not to be overwritten by fastplong. In this case, fastplong will report an error and quit if it finds any of the output files (read, json report, html report) already exists before.
split the output to multiple files for parallel processing
See output splitting
fastplong workflow
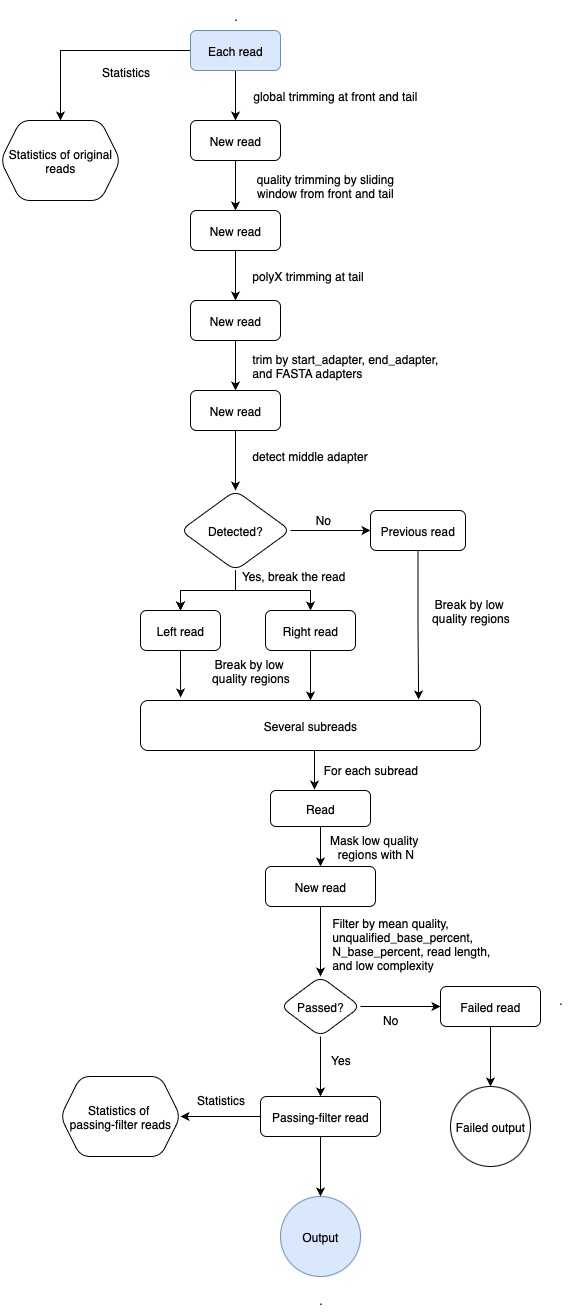
filtering
Multiple filters have been implemented.
quality filter
Quality filtering is enabled by default, but you can disable it by -Q or disable_quality_filtering.
fastplong supports filtering by limiting the N base number (--n_base_limit, disabled by default) and N base percentage (-n, --n_percent_limit, enabled by default). For example, to limit the N base no more than 100, and no more than 20%, you can use the command:
fastplong -i in.fq -o out.fq --n_base_limit 100 --n_percent_limit 20
To filter reads by its percentage of unqualified bases, two options should be provided:
-q, --qualified_quality_phredthe quality value that a base is qualified. Default 15 means phred quality >=Q15 is qualified.-u, --unqualified_percent_limithow many percents of bases are allowed to be unqualified (0~100). Default 40 means 40%
You can also filter reads by its average quality score
-m, --mean_qualif one read's average quality score <avg_qual, then this read is discarded. Default 0 means no requirement (int [=0])
length filter
Length filtering is enabled by default, but you can disable it by -L or --disable_length_filtering. The minimum length requirement is specified with -l or --length_required.
You can specify --length_limit to discard the reads longer than length_limit. The default value 0 means no limitation.
Other filter
New filters are being implemented. If you have a new idea or new request, please file an issue.
adapters
fastplong trims adapter in both read start and read end. Adapter trimming is enabled by default, but you can disable it by -A or --disable_adapter_trimming.
fastplong -i in.fq -o out.fq -s AAGGATTCATTCCCACGGTAACAC -e GTGTTACCGTGGGAATGAATCCTT
-
If the adapter sequences are known, it's recommended to specify
-s, --start_adapterfor read start adapter sequence, and-e, --end_adapterfor read end adapter sequence as well. -
If
--end_adapteris not specified but--start_adapteris specified, then fastplong will use the reverse complement sequence ofstart_adapterto beend_adapter. -
You can also specify
-a, --adapter_fastato give a FASTA file to tellfastplongto trim multiple adapters in this FASTA file. Here is a sample of such adapter FASTA file:
>Adapter 1
AGATCGGAAGAGCACACGTCTGAACTCCAGTCA
>Adapter 2
AGATCGGAAGAGCGTCGTGTAGGGAAAGAGTGT
>polyA
AAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAA
-
The adapter sequence in the FASTA file should be at least 6bp long, otherwise it will be skipped. And you can give whatever you want to trim, rather than regular sequencing adapters (i.e. polyA).
-
If all these adapter options (
start_adapter,end_adapterandadapter_fasta) are not specified,fastplongwill try to detect the read start and read end adapters automatically. The detected adapter sequences may be a bit shorter or longer than the real ones. And there is a certain probability of misidentification, especially when most reads don't have adapters (it won't cause too bad result in this case). -
fastplong calculates edit distance when detecting adapters. You can specify the
-d, --distance_thresholdto adjust the mismatch tolerance of adapter comparing. The default value is 0.25, which means allowing 25% mismatch ratio (i.e. allow 10 distance for 40bp adapter). Suggest to increase this value when the data is much noisy (high error rate), and decrease this value when the data is with high quality (low error rate). -
to make a cleaner trimming, fastplong will trim a little more bases connected to the adapters. This option can be specified by
--trimming_extension, with a default value of 10.
per read cutting by quality score
fastplong supports per read sliding window cutting by evaluating the mean quality scores in the sliding window. fastplong supports 2 different operations, and you enable one or both:
-5, --cut_frontmove a sliding window from front (5') to tail, drop the bases in the window if its mean quality is below cut_mean_quality, stop otherwise. Default is disabled. The leading N bases are also trimmed. Usecut_front_window_sizeto set the widnow size, a
Related Skills
node-connect
344.4kDiagnose OpenClaw node connection and pairing failures for Android, iOS, and macOS companion apps
frontend-design
99.2kCreate distinctive, production-grade frontend interfaces with high design quality. Use this skill when the user asks to build web components, pages, or applications. Generates creative, polished code that avoids generic AI aesthetics.
openai-whisper-api
344.4kTranscribe audio via OpenAI Audio Transcriptions API (Whisper).
qqbot-media
344.4kQQBot 富媒体收发能力。使用 <qqmedia> 标签,系统根据文件扩展名自动识别类型(图片/语音/视频/文件)。
