MedTator
A Serverless Text Annotation Tool for Corpus Development
Install / Use
/learn @OHNLP/MedTatorREADME
<img alt="MedTator" src="https://raw.githubusercontent.com/wiki/OHNLP/MedTator/img/logo.png">
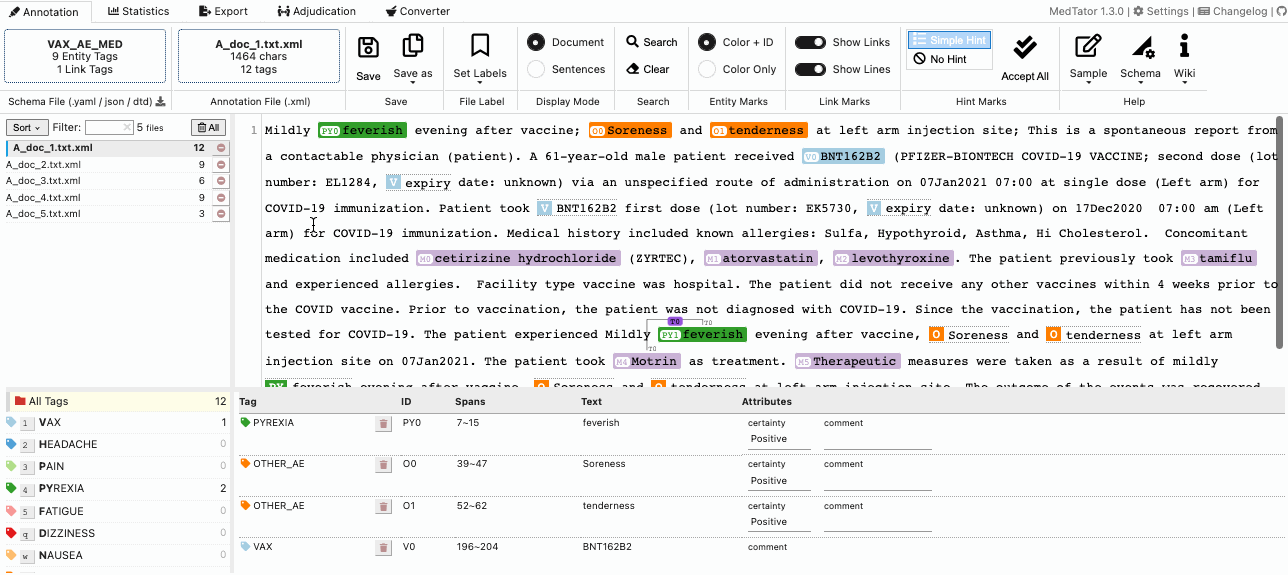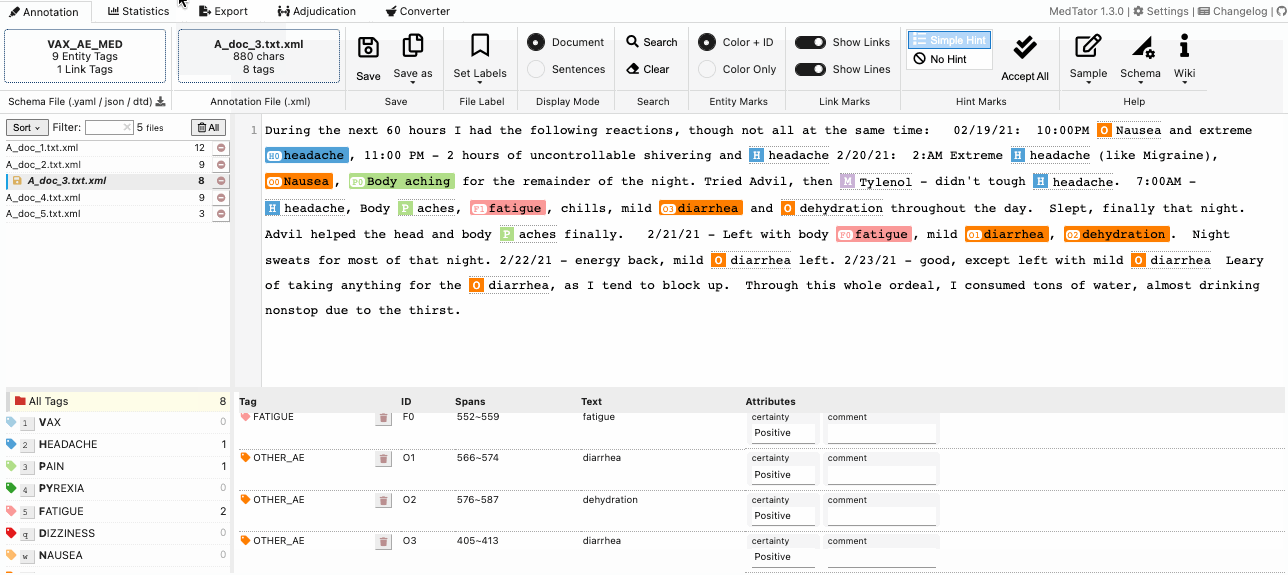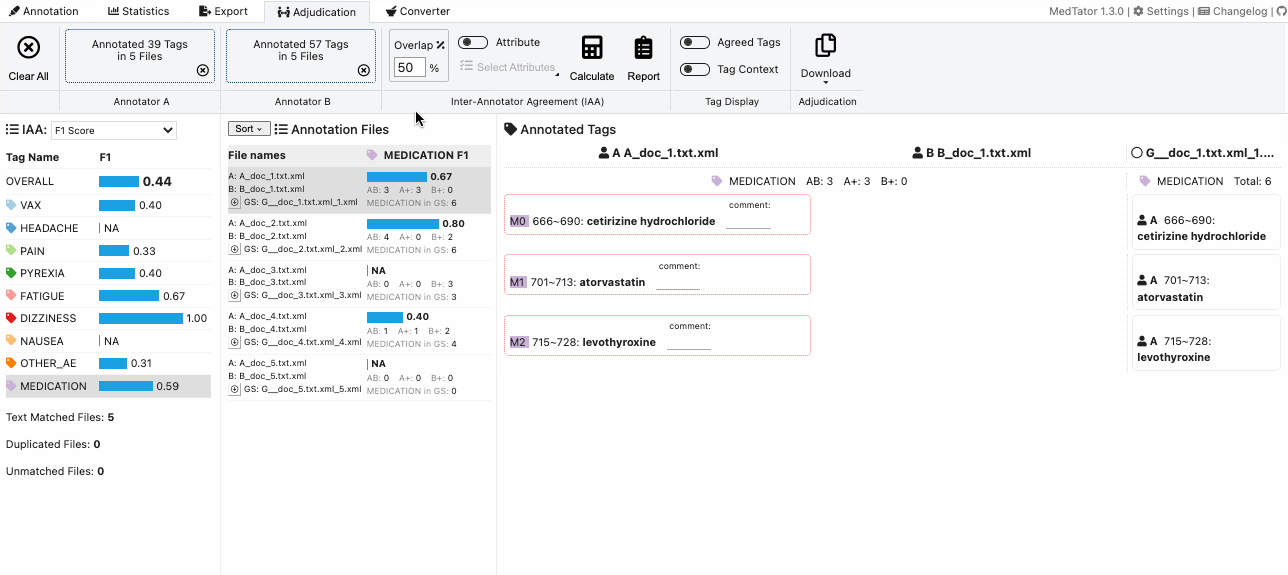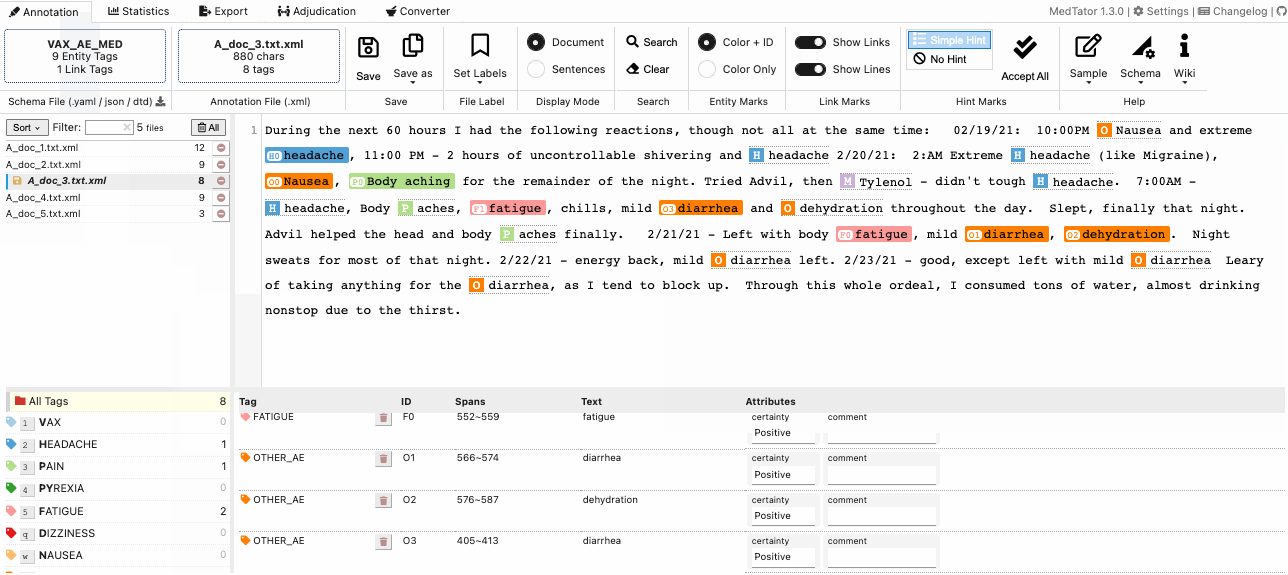
MedTator is a serverless text annotation tool for corpus development. It is built on HTML5 techniques and many open-source packages, and was designed to be easy-to-use for your annotation task.
No Java, no Python, no PHP, no Docker, no MySQL, and no need to install any server or client runtime for corpus annotation! Check Here to Start Annotation Now!
If you're having trouble using MedTator, you can use Issues to tell us about the issue you're experiencing.
Documentation
- New to MedTator? Let's have a Quick Start.
- More samples? Check our Sample Datasets.
- No internet access? Check this Standalone version
- Effective strategies for annotating? Check this Annotation best practices
- Design annotation schema? Check this Annotation Schema Design.
- Parse annotation data? Here is the Annotation File Format.
- Use annotated files for machine learning? We have some Python Sample Scripts.
- Having issues? Check the Issues.
- More details about MedTator? We have a Manual.
- And more contents in the Wiki
MedTator Development
MedTator itself doesn't require Python runtime environment, so you don't need to install any runtime environment to run MedTator for corpus annotation. If you are interested in the MedTator development or just want to try the development version, a Python 3+ runtime environment is needed to run a debugging server.
You can install a Python 3+ or Miniconda / Anaconda, then download the source code of MedTator and install the requirements (just Python Flask, that's all):
pip install -r requirements.txt
Then, run the following command to start a local server which is binding port 8086:
python web.py
Now you can open web browser and check the http://localhost:8086/.
For more details of the parameters for web.py, run python web.py -h and it will show the details as follows.
usage: web.py [-h] [--mode {build,run,release}] [--lib {local,cdn}]
[--path PATH] [--fn FN]
MedTator Development Server and Toolkit
optional arguments:
-h, --help show this help message and exit
--mode {build,run,release}
What do you want to do? `run` for starting the
development server. `build` for generating a static
HTML page for public release or local release.
--lib {local,cdn} Where to get third party libs in the HTML page? If
choose local, please make sure to copy the `static`
folder after generated the HTML file.
--path PATH Which folder to be used for the output page? The
default folder is the docs/ folder for public release.
--fn FN What file name to be used for the output page? The
default file name is the `index.html` which could be
accessed directly by browser.
Build the static version
To update the static version for publication (e.g., GitHub Pages), run the following command. It will generate a static HTML file in the docs/ folder and copy other files.
In addition, as the default filename is index.html, the build script will automatically create a index.VERSION.html in the build path for backup.
So that the user can access the old version for comparison or checking old functions.
python web.py --mode build
Or you can build a dev version for public testing, run the following command:
python web.py --mode build --fn dev.html
Or you can build a standalone version for local use, run the following command:
python web.py --mode build --lib local --fn standalone.html
Then, you can create a release zip file:
python web.py --mode release
License
Apache-2.0 License
Citation
If you use MedTator in scientific work or want to learn more about it, please take a look at our paper:
He H, Fu S, Wang L, Liu S, Wen A, Liu H. MedTator: a serverless annotation tool for corpus development. Bioinformatics, Volume 38, Issue 6, 15 March 2022, Pages 1776–1778, DOI: 10.1093/bioinformatics/btab880, PMID: 34983060
Change log
1.3.x (2023-05)
- Research on IOB2/BIO format editing
- Research on UMAP algorithm
- Research on in-browser text embedding
- Add sample dataset for error analysis
- Update icon for error analysis
- Update Cohen's Kappa calculation
- Update README for new features
- Update documents for error analysis
- Update documents for new schema
- Update UI for error analysis
- Update demo script for inter-sentence
1.3.16 (2023-04-03)
- Added example Python script for masking entities
- Updated medtator_kits.py for saving xml
- Updated relation annotation example schema and data
- Fixed one-offset bug in first-line newline sign
- Fixed cross-line entity render bug
1.3.15 (2023-02-26)
- Added workspace JSON drag and drop on file list
- Added workspace saving function
- Added workspace loading function
- Added schema drag and drop on file list
- Added shortcut ALT + ↑ / ↓ to move to prev / next file
- Added Toolkit/MedTaggerVis for checking .ann files (experimental)
- Added local setting cache for auto save/load (experimental)
- Fixed error analysis Sankey link bug
- Fixed error analysis SVG height bug
- Fixed error analysis popup box bug
- Updated design for .ann/.txt conversion
- Updated FAQ in MedTator Wiki
- Updated scripts for downstream tasks
- Updated sample datasets for new settings
1.3.11 (2022-12-08)
- Added annotation visualization based on brat
- Added Python scripts for read/parse MedTator XML format
- Added Jupyter notebook for downstream tasks with MedTator XML files.
- Developed functions for brat vis format conversion
- Developed tag extraction based offset spans
- Updated style for link/relation display
1.3.8 (2022-10-27)
- Added schema of .yml extension support
- Added auto-save (experimental) feature
- Fixed scheme editor open bug
1.3.7 (2022-09-29)
- Update the UI for IAA
1.3.6 (2022-09-15)
- Updated a data sample for error analysis
1.3.5 (2022-08-18)
- Added label panel for error labeling
- Added error sankey diagram for error analysis
- Added token scatter for error analysis
- Added heatmap of error distribution for error analysis
- Added bar charts of error statistics for error analysis
- Added tag list and menu items for error analysis
- Updated scripts for text embedding web services
- Fixed IOB2/BIO empty export bug
- Fixed exporter window height bug
- Fixed BioC format export annotation text encoding bug
- Fixed BioC format export attribute text encoding bug
1.3.2 (2022-08-04)
- Added functions for error analysis
- Added Math.js for statistics and matrix
- Added ECharts for visualization
- Added a sample dataset for error analysis
- Updated the UI design for adjudication tab
- Updated the packaged used to reduce loading time
- Fixed open annotation files dialog bug
- Fixed IAA download bug
1.3.0 (2022-07-21)
- Refactored file reading workflow with asynchronous processing
- Refactored annotation file loading workflow
- Refactored code structure to reduce code size
- Designed JSON/YAML format schema
- Added tokenization exceptions
- Added pagination for large dataset
- Added loading dataset annimation
- Added drag and drop folder for adjudication
- Added drag and drop folder for converter
- Added converting text files to MedTator xml
- Added a float text content viewer
- Added reading JSON/YAML for schema editor
- Added JSON/YAML download option for schema editor
- Added cmd/ctrl + s shortcut key for saving current file
- Updated annotation UI design for bigger drag&drop area
- Updated converter UI design for bigger buttons
- Updated converter for downloading individual result file
- Updated converter with support of viewing contents
- Updated event handlers for drag and drop events
- Updated license files of imported libraries
- Updated the sample schemas with JSON/YAML format
1.2.48 (2022-07-07)
- Added function for customizing sample text
- Added changelog information on UI
- Added sorting for corpus token list in statistics
- Added tag details for corpus summary in statistics
- Added filter options for token statistics
- Added filtered count in token summaryies
- Added a remote corpus sample
- Updated all sample datasets
- Updated the visual design for corpus summary in statistics
- Updated the color encoding for annotated tags in statistics
- Updated scripts for experimental tasks
- Fixed ribbon menu resize bug
1.2.41 (2022-06-23)
- Added meta-data structure for annotation file
- Added color label to annotation file
- Added functions for updating annotator for adjudication
- Added download all tags for adjudication
- Added sorting by label color
- Added MedTagger ann format converter
- Added UI for converting MedTagger results
- Added folder drag an
