Masif
MaSIF- Molecular surface interaction fingerprints. Geometric deep learning to decipher patterns in molecular surfaces.
Install / Use
/learn @LPDI-EPFL/MasifREADME
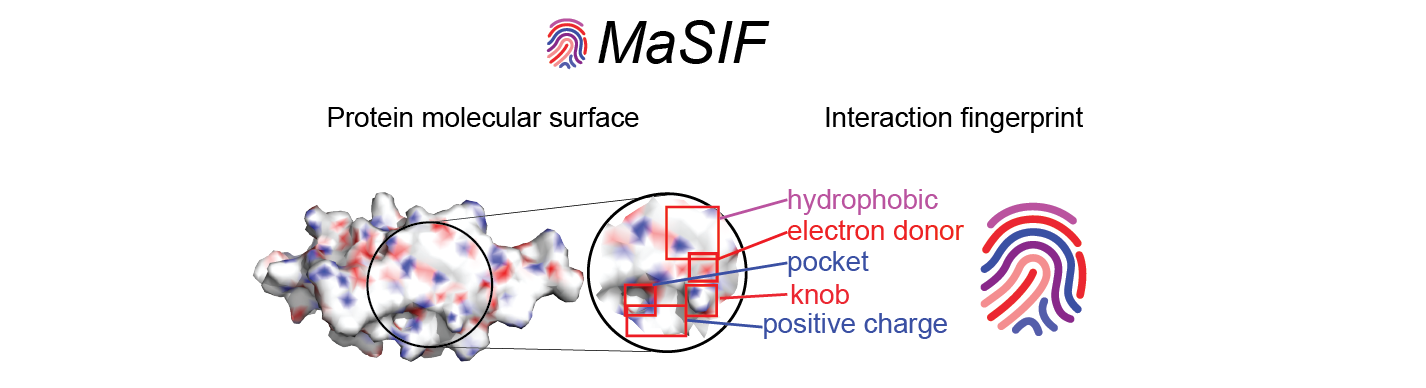
MaSIF- Molecular Surface Interaction Fingerprints: Geometric deep learning to decipher patterns in protein molecular surfaces.
Table of Contents:
- Description
- System and hardware requirements
- Software prerequisites
- Installation
- Method overview
- MaSIF applications
- PyMOL plugin
- Docker container
- License
- Reference
Description
MaSIF is a proof-of-concept method to decipher patterns in protein surfaces important for specific biomolecular interactions. To achieve this, MaSIF exploits techniques from the field of geometric deep learning. First, MaSIF decomposes a surface into overlapping radial patches with a fixed geodesic radius, wherein each point is assigned an array of geometric and chemical features. MaSIF then computes a descriptor for each surface patch, a vector that encodes a description of the features present in the patch. Then, this descriptor can be processed in a set of additional layers where different interactions can be classified. The features encoded in each descriptor and the final output depend on the application-specific training data and the optimization objective, meaning that the same architecture can be repurposed for various tasks.
This repository contains a protocol to prepare protein structure files into feature-rich surfaces (with both geometric and chemical features), to decompose these into patches, and tensorflow-based neural network code to identify patterns in these using deep geometric learning. To show the potential of the approach, we showcase three proof-of-concept applications: a) ligand prediction for protein binding pockets (MaSIF-ligand); b) protein-protein interaction (PPI) site prediction in protein surfaces, to predict which surface patches on a protein are more likely to interact with other proteins (MaSIF-site); c) ultrafast scanning of surfaces, where we use surface fingerprints from binding partners to predict the structural configuration of protein-protein complexes (MaSIF-search).
This repository should closely reproduce the experiments of:
Gainza, P., Sverrisson, F., Monti, F., Rodola, E., Boscaini, D Bronstein, M. M., & Correia, B. E. (2019). Deciphering interaction fingerprints from protein molecular surfaces using geometric deep learning. Nat Methods 17, 184–192 (2020). https://doi.org/10.1038/s41592-019-0666-6
<span style="color:red">Note: Since Feb 2020, we have greatly simplified the installation of MaSIF by replacing all Matlab code with Python code. However, this slightly changes the results from the paper. To reproduce the results for the paper exactly as published (with the pretrained neural networks) you can obtain it at: https://github.com/pablogainza/masif_paper </span>.
MaSIF is distributed under an Apache License. This code is meant to serve as a tutorial, and the basis for researchers to exploit MaSIF in protein-surface learning tasks.
System and hardware requirements
MaSIF has been tested on both Linux (Red Hat Enterprise Linux Server release 7.4, with a Intel(R) Xeon(R) CPU E5-2650 v2 @ 2.60GHz processesor and 16GB of memory allotment) and Mac OS environments (macOS High Sierra, processor 2.8 GHz Intel Core i7, 16GB memory). To reproduce the experiments in the paper, the entire datasets for all proteins consume about 1.4 terabytes.
Currently, MaSIF takes about 2 minutes to preprocess every protein. For this reason, we recommend a distributed cluster to preprocess the data for large datasets of proteins. Once data has been preprocessed, we strongly recommend using a GPU to train or evaluate the trained models as it can be up to 100 times faster than a CPU.
Software prerequisites
MaSIF relies on external software/libraries to handle protein databank files and surface files, to compute chemical/geometric features and coordinates, and to perform neural network calculations. The following is the list of required libraries and programs, as well as the version on which it was tested (in parenthesis).
- Python (3.6)
- reduce (3.23). To add protons to proteins.
- MSMS (2.6.1). To compute the surface of proteins.
- BioPython (1.66) . To parse PDB files.
- PyMesh (0.1.14). To handle ply surface files, attributes, and to regularize meshes.
- PDB2PQR (2.1.1), multivalue, and APBS (1.5). These programs are necessary to compute electrostatics charges.
- open3D (0.5.0.0). Mainly used for RANSAC alignment.
- Tensorflow (1.9). Use to model, train, and evaluate the actual neural networks. Models were trained and evaluated on a NVIDIA Tesla K40 GPU.
- StrBioInfo. Used for parsing PDB files and generate biological assembly for MaSIF-ligand.
- Dask (2.2.0). Run function calls on multiple threads (Optional for reproducing some benchmarks).
- Pymol. This optional plugin allows one to visualize surface files in PyMOL.
Alternatively you can use the Docker version, which is the easiest to install (See Docker container)
Installation
After preinstalling dependencies, add the following environment variables to your path, changing the appropriate directories:
export APBS_BIN=/path/to/apbs/APBS-1.5-linux64/bin/apbs
export MULTIVALUE_BIN=/path/to/apbs/APBS-1.5-linux64/share/apbs/tools/bin/multivalue
export PDB2PQR_BIN=/path/to/apbs/apbs/pdb2pqr-linux-bin64-2.1.1/pdb2pqr
export PATH=$PATH:/path/to/reduce/
export REDUCE_HET_DICT=/path/to/reduce/reduce_wwPDB_het_dict.txt
export PYMESH_PATH=/path/to/PyMesh
export MSMS_BIN=/path/to/msms/msms
export PDB2XYZRN=/path/to/msms/pdb_to_xyzrn
Clone masif to a local directory
git clone https://github.com/lpdi-epfl/masif
cd masif/
Since MaSIF is written in Python, no compilation is required.
Method overview
From a protein structure MaSIF computes a molecular surface discretized as a mesh according to the solvent excluded surface (computed using MSMS), and assigns geometric and chemical features to every point (vertex) in the mesh. Around each vertex of the mesh, we extract a patch with geodesic radius of r=9 Å or r=12 Å. Then, MaSIF applies a geometric deep neural network to these patches. The neural network consists of one or more layers applied sequentially; a key component of the architecture is the geodesic convolution, generalizing the classical convolution to surfaces and implemented as an operation on local patches.
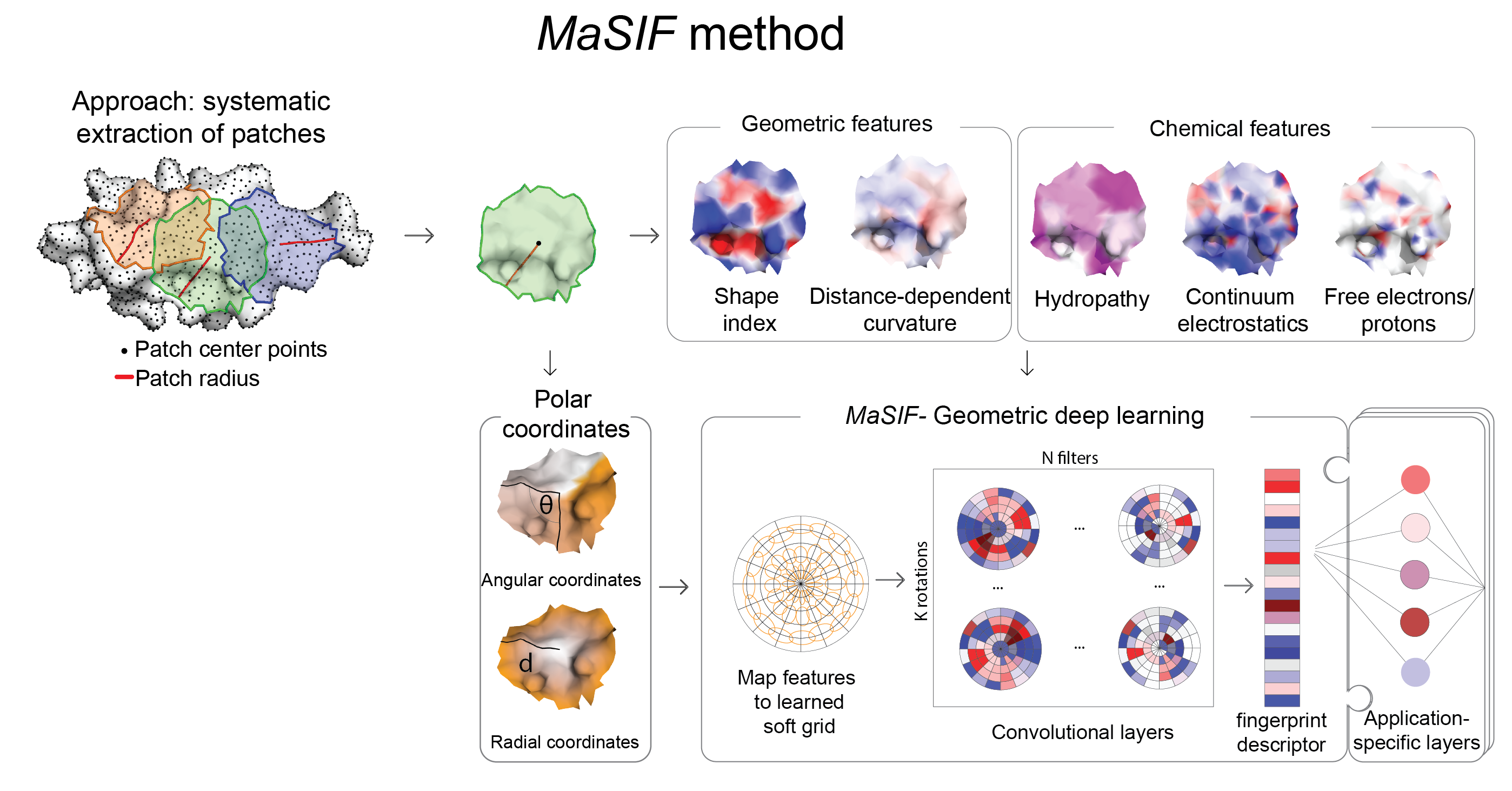
The procedure is repeated for different patch locations similarly to a sliding window operation on images, producing the surface fingerprint descriptor at each point, in the form of a vector that stores information about the surface patterns of the center point and its neighborhood. The parameter set minimizes a cost function on the training dataset, which is specific to each application that we present here.
MaSIF data preparation
For each application, MaSIF requires a preprocessing of data. This entails a running a scripted protocol, which performs the following steps:
- Download the PDB.
- Protonate the PDB, extract the desired chains, triangulate the surface (using MSMS), and compute chemical features.
- Extract all patches, with features and coordinates, for each protein.
MaSIF's main speed bottleneck lie in these three steps. The main performance bottlenecks are computing the angular coordinates using MDS, computing the Poisson-Boltzmann electrostatics and regularizing the mesh after computing the MSMS surface.
Each application data directory (under masif/data/masif*) contains a script to precompute the data.
To run this protocol for a single protein, (e.g. chain A of PDB id code 1MBN ) run:
./data_prepare_one.sh 1MBN_A_
To run it on a pair of interacting protein domains (chains A,B, of PDB id 1AKJ form the first domain and chains D,E form the second domain):
./data_prepare_one.sh 1AKJ_AB_DE
If you have access to a cluster (strongly recommended), then this process can be run in parallel. If your cluster supports slurm files, we provide a slurm file under each application data directory. which can be run using sbatch:
sbatch data_prepare.slurm
Most of the PDBs that were used for the paper, and their corresponding surfaces (with precomputed chemical features) are available at: https://doi.org/10.5281/zenodo.2625420 . The unbound proteins are available in this repository under data/masif_ppi_search_ub/data_preparation/00-raw_pdbs/.
Note that the preparation of the data can consume a large amount of space for large protein databases. This is due to the fact that the preprocessing step decomposes protein surfaces into overlapping patche

