BiaPy
Open source Python library for building bioimage analysis pipelines
Install / Use
/learn @BiaPyX/BiaPyREADME

BiaPy: Accessible deep learning on bioimages
<p align="left"> <a href="https://www.python.org/"> <img src="https://img.shields.io/badge/Python-3.11-yellow.svg" /></a> <a href= "https://pytorch.org/"> <img src="https://img.shields.io/badge/PyTorch-2.9.1-orange.svg" /></a> <a href= "https://pypi.org/project/biapy/"> <img src="https://img.shields.io/pypi/v/biapy.svg?color=green" /></a> <a href= "https://anaconda.org/conda-forge/biapy/"> <img src="https://anaconda.org/conda-forge/biapy/badges/version.svg" /></a> <a href= "https://github.com/BiaPyX/BiaPy/blob/master/LICENSE"> <img src="https://img.shields.io/badge/License-MIT-blue.svg" /></a> <a href= "https://biapy.readthedocs.io/en/latest/"> <img src="https://img.shields.io/badge/Doc-Latest-2BAF2B.svg" /></a> <a href= "https://www.nature.com/articles/s41592-025-02699-y"> <img src="https://img.shields.io/badge/Nature_Methods-Paper-bd2635.svg" /></a> <a href= "https://ieeexplore.ieee.org/abstract/document/10230593"> <img src="https://img.shields.io/badge/IEEE-Paper-00629B.svg" /></a> </p>🔥NEWS🔥: BiaPy's paper is finally out in Nature Methods! Check it out at: https://www.nature.com/articles/s41592-025-02699-y
For those who have no access to Nature Methods, here is the preprint at bioRxiv: https://www.biorxiv.org/content/10.1101/2024.02.03.576026v3
BiaPy is an open source library and application that streamlines the use of common deep-learning workflows for a large variety of bioimage analysis tasks, including 2D and 3D semantic segmentation, instance segmentation, object detection, image denoising, single image super-resolution, self-supervised learning, image classification and image to image translation.
BiaPy is a versatile platform designed to accommodate both proficient computer scientists and users less experienced in programming. It offers diverse and user-friendly access points to our workflows.
This repository is actively under development by the BiaPy Team, a group of contributors from different research institutions.
Description videos
Find a comprehensive overview of BiaPy and its functionality in the following videos:
|  <br> <span style="font-weight:normal">BiaPy history and GUI demo at RTmfm by Ignacio Arganda-Carreras and Daniel Franco-Barranco.</span> |
|-------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------|
|
<br> <span style="font-weight:normal">BiaPy history and GUI demo at RTmfm by Ignacio Arganda-Carreras and Daniel Franco-Barranco.</span> |
|-------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------|
|  <br> BiaPy presentation at Virtual Pub of Euro-BioImaging by Ignacio Arganda-Carreras. | | | | |
<br> BiaPy presentation at Virtual Pub of Euro-BioImaging by Ignacio Arganda-Carreras. | | | | |
User interface
You can also use BiaPy through our graphical user interface (GUI).
Download BiaPy GUI for you OS
Project's page: [BiaPy GUI]
Scientific publications using BiaPy
| | |
|--------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------|:-----------------------------------------------------------------------------------------------------------------------------------------------------------:|
| López-Cano, Daniel, et al. "Characterizing Structure Formation through Instance Segmentation" (2023). <br><br> This study presents a machine-learning framework to predict the formation of dark matter haloes from early universe density perturbations. Utilizing two neural networks, it distinguishes particles comprising haloes and groups them by membership. The framework accurately predicts halo masses and shapes, and compares favorably with N-body simulations. The open-source model could enhance analytical methods of structure formation by analyzing initial condition variations. BiaPy is used in the creation of the watershed approach. <br><br> [Documentation (not yet)] [Paper] |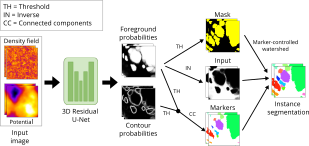  |
|
|
| Franco-Barranco, Daniel, et al. "Current Progress and Challenges in Large-scale 3D Mitochondria Instance Segmentation." (2023). <br><br> This paper reports the results of the MitoEM challenge on 3D instance segmentation of mitochondria in electron microscopy images, held in conjunction with IEEE-ISBI 2021. The paper discusses the top-performing methods, addresses ground truth errors, and proposes a new scoring system to improve segmentation evaluation. Despite progress, challenges remain in segmenting mitochondria with complex shapes, keeping the competiti

